 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_36240 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 717aa MW: 78786 Da PI: 8.0043 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 76.6 | 2.7e-24 | 116 | 217 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefel 92
f+k+lt sd++++g +++p++ ae++ + ++ s++l+ +d++g +W++++iyr++++r++lt+GW+ Fv+ ++L +gD v+F + ++el
PEQU_36240 116 FCKTLTASDTSTHGGFSVPRRAAEDCfppldyrQ--QRPSQELIAKDLHGVEWRFRHIYRGQPRRHLLTTGWSAFVNKKKLVSGDAVLFLR--GGNGEL 210
99****************************9743..345679************************************************4..378889 PP
EEEEE-S CS
B3 93 vvkvfrk 99
+++++r+
PEQU_36240 211 RLGIRRA 217
****997 PP
| |||||||
| 2 | Auxin_resp | 114.8 | 7.5e-38 | 245 | 325 | 2 | 82 |
Auxin_resp 2 ahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsL 82
a+a+stk++F + YnPra +seF+v+++k++k+++ ++svG+Rfkm+fe+ed++err++G++++vsdldp++Wp+SkWr+L
PEQU_36240 245 ANAMSTKKMFPINYNPRANQSEFIVPYWKFRKSCSYSISVGTRFKMRFESEDAAERRCTGLITAVSDLDPIQWPGSKWRCL 325
789*****************************************************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 4.4E-41 | 108 | 222 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.19E-42 | 113 | 241 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.64E-23 | 114 | 216 | No hit | No description |
| Pfam | PF02362 | 3.4E-22 | 116 | 217 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.592 | 116 | 218 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 5.6E-25 | 116 | 218 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.5E-32 | 245 | 325 | IPR010525 | Auxin response factor |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009725 | Biological Process | response to hormone | ||||
| GO:0010050 | Biological Process | vegetative phase change | ||||
| GO:0010158 | Biological Process | abaxial cell fate specification | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 717 aa Download sequence Send to blast |
MCLELWHACA GPRISLPRKG SLVVYLPQGH LEHLAGGQRS AGGGMLIRYD VPPHVFCRVV 60 EVQLRVHAAT DEVYAQLSMI AENEAFEKQI QGGVEGEGEE VDEMDGGDQS PVPHMFCKTL 120 TASDTSTHGG FSVPRRAAED CFPPLDYRQQ RPSQELIAKD LHGVEWRFRH IYRGQPRRHL 180 LTTGWSAFVN KKKLVSGDAV LFLRGGNGEL RLGIRRASQI KGNIAHNISG SQNLNVGPGS 240 LASLANAMST KKMFPINYNP RANQSEFIVP YWKFRKSCSY SISVGTRFKM RFESEDAAER 300 RCTGLITAVS DLDPIQWPGS KWRCLVVRWD EDDVDNSLHN RVSPWEVEPT GSVSCPNSLL 360 ASGSKRTRIS IPSISADYPH PSGSGYQSLR ESARFQEVLQ GQEIFGFKSP YNGGDATLPH 420 LSEMRSHHLP EMRRCIANPN SCSLAGSGGN ARVPPGNSDF HFQGTGFGES MRFHKVLQGQ 480 EIVPVLPSFR GLAVESHREN DGSKIFESYS AGRCSAPPLQ GYCTFIQPCT TAAQVSSPSS 540 VLMFQQAASQ LPCTYSLYDT NEKEKADDSN LSAPISSAEG TRVQPASFTT RCENLTRFKA 600 PVHKNGQSII HGSGSSCRLF GFPLTEKIPV TNVVASSLNE SPLIEPSIKS SFPSQRPQIP 660 TNNPGHGCTR VSVFYLSVAT FWNSLDFSSL LHHEGFSVPY LLQAGLLISR SLLDMPI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-109 | 1 | 354 | 19 | 359 | Auxin response factor 1 |
| 4ldw_A | 1e-109 | 1 | 354 | 19 | 359 | Auxin response factor 1 |
| 4ldw_B | 1e-109 | 1 | 354 | 19 | 359 | Auxin response factor 1 |
| 4ldx_A | 1e-109 | 1 | 354 | 19 | 359 | Auxin response factor 1 |
| 4ldx_B | 1e-109 | 1 | 354 | 19 | 359 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00021 | PBM | Transfer from AT2G33860 | Download |
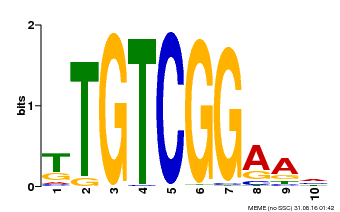
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020595811.1 | 0.0 | auxin response factor 3-like isoform X2 | ||||
| Refseq | XP_020595812.1 | 0.0 | auxin response factor 3-like isoform X2 | ||||
| TrEMBL | A0A2I0AU81 | 0.0 | A0A2I0AU81_9ASPA; Auxin response factor | ||||
| STRING | XP_008798266.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2537 | 35 | 80 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33860.1 | 1e-177 | ARF family protein | ||||



