 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_09738 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 375aa MW: 41847.6 Da PI: 8.6607 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 85.7 | 4.6e-27 | 101 | 155 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFveaveqL G +kA+P++il +m+++ Lt+++v+SHLQkYR++
PEQU_09738 101 KVKVDWTPELHRRFVEAVEQL-GVDKAVPSRILVIMGIQNLTRHNVASHLQKYRSH 155
5689*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.057 | 98 | 157 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-26 | 99 | 158 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 9.86E-18 | 99 | 158 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 8.1E-25 | 101 | 155 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.5E-8 | 104 | 153 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0010638 | Biological Process | positive regulation of organelle organization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 375 aa Download sequence Send to blast |
MLALPPLINS KAEEKEGFSD AIPGDDILDS INFSCIFENF DVDGDLFPTE LQFSEPQHIL 60 PEFSAEGEEQ ENSSQSQKRE EKKDELSATG KTVKRSHGKK KVKVDWTPEL HRRFVEAVEQ 120 LGVDKAVPSR ILVIMGIQNL TRHNVASHLQ KYRSHRKHLV AREAEAASWT QRRQLYASAA 180 GRRNTRSIIM KKENSTRIAP TIGFPPPPPP FHPQFRPLHV WGHPTTGRRP MFQAMWTLGH 240 AATPPWPPHL PNYIPYLQPN YRLLQGAAEG WGTSAVMSTP LTPFQLPAIR FAPPPVAGVL 300 AHPTYGSQHL VSPLKQHSNL QPEHSDYPSK ESVDAAIGDV LAKPWLPLPL GLKPPSTDSV 360 LLELQRQLGC NPDGN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 1e-14 | 98 | 153 | 2 | 57 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that promotes chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport. {ECO:0000269|PubMed:19808806}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
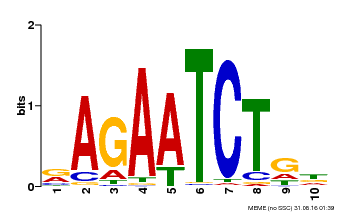
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. {ECO:0000269|PubMed:11340194}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020575235.1 | 0.0 | probable transcription factor GLK1 isoform X1 | ||||
| Swissprot | Q5Z5I4 | 3e-53 | GLK1_ORYSJ; Probable transcription factor GLK1 | ||||
| TrEMBL | A0A2I0WC58 | 1e-147 | A0A2I0WC58_9ASPA; Transcription activator GLK1 | ||||
| STRING | XP_010260326.1 | 2e-85 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2415 | 38 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 4e-56 | GBF's pro-rich region-interacting factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



