 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_06885 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | LBD | ||||||||
| Protein Properties | Length: 126aa MW: 14043.1 Da PI: 8.7499 | ||||||||
| Description | LBD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF260 | 131.2 | 4.4e-41 | 21 | 119 | 1 | 100 |
DUF260 1 aCaaCkvlrrkCakdCvlapyfpaeqpkkfanvhklFGasnvlkllkalpeeeredamsslvyeAearardPvyGavgvilklqqqleqlkaelallke 99
+Ca+Ck+lrrkC++dCv+apyfp +qp kf nvh++FGasnv+k+l++l++ ereda++sl+yeAe+r+rdPvyG++g+ + lq+ql++l+ +l+++++
PEQU_06885 21 PCAGCKFLRRKCQPDCVFAPYFPPDQPTKFSNVHRVFGASNVTKILNDLKPFEREDAVNSLAYEAEMRLRDPVYGCAGI-SLLQRQLRELQIDLSRARS 118
7****************************************************************************95.78*************9988 PP
DUF260 100 e 100
e
PEQU_06885 119 E 119
7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50891 | 24.245 | 20 | 120 | IPR004883 | Lateral organ boundaries, LOB |
| Pfam | PF03195 | 3.5E-40 | 21 | 117 | IPR004883 | Lateral organ boundaries, LOB |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009799 | Biological Process | specification of symmetry | ||||
| GO:0009944 | Biological Process | polarity specification of adaxial/abaxial axis | ||||
| GO:0009954 | Biological Process | proximal/distal pattern formation | ||||
| GO:0048441 | Biological Process | petal development | ||||
| GO:0005654 | Cellular Component | nucleoplasm | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 126 aa Download sequence Send to blast |
NVLTIPTPPT MASGGHTVSS PCAGCKFLRR KCQPDCVFAP YFPPDQPTKF SNVHRVFGAS 60 NVTKILNDLK PFEREDAVNS LAYEAEMRLR DPVYGCAGIS LLQRQLRELQ IDLSRARSEL 120 VKYQSA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5ly0_A | 2e-51 | 9 | 123 | 3 | 114 | LOB family transfactor Ramosa2.1 |
| 5ly0_B | 2e-51 | 9 | 123 | 3 | 114 | LOB family transfactor Ramosa2.1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Promotes the switch from proliferation to differentiation in the embryo sac. Negative regulator of cell proliferation in the adaxial side of leaves. Regulates the formation of a symmetric lamina and the establishment of venation. Interacts directly with RS2 (rough sheath 2) to repress some knox homeobox genes. {ECO:0000269|PubMed:17209126}. | |||||
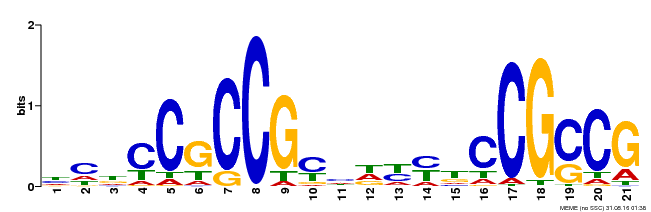
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00211 | DAP | Transfer from AT1G65620 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020702209.1 | 5e-66 | protein ASYMMETRIC LEAVES 2 | ||||
| Refseq | XP_020702210.1 | 5e-66 | protein ASYMMETRIC LEAVES 2 | ||||
| Refseq | XP_020702211.1 | 5e-66 | protein ASYMMETRIC LEAVES 2 | ||||
| Swissprot | Q32SG3 | 6e-59 | LBD6_MAIZE; LOB domain-containing protein 6 | ||||
| TrEMBL | A0A2I0VVY1 | 1e-64 | A0A2I0VVY1_9ASPA; LOB domain-containing protein 6 | ||||
| STRING | VIT_00s0340g00090.t01 | 3e-58 | (Vitis vinifera) | ||||
| STRING | Pavir.J03307.1.p | 5e-58 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2267 | 38 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G65620.4 | 7e-56 | LBD family protein | ||||



