 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s1210051g004 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1090aa MW: 123005 Da PI: 5.8391 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 143.4 | 6.2e-45 | 64 | 151 | 31 | 118 |
CG-1 31 ktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlLeeelekivlvhylevk 118
+ r gsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+een++fqrr+yw+Lee++ +ivlvhylevk
PDK_30s1210051g004 64 RGRSAGGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKAGSIDVLHCYYAHGEENENFQRRSYWMLEEDYMHIVLVHYLEVK 151
567789*******************************************************************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 64.214 | 42 | 156 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.1E-53 | 46 | 151 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.8E-37 | 60 | 150 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 1.0E-5 | 478 | 568 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.6E-7 | 481 | 559 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.97E-17 | 481 | 567 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 2.4E-18 | 663 | 778 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.78E-15 | 672 | 775 | No hit | No description |
| SuperFamily | SSF48403 | 2.49E-17 | 673 | 778 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.555 | 674 | 777 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 2.0E-7 | 688 | 778 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.028 | 716 | 745 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 560 | 755 | 784 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 7.7E-9 | 888 | 944 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.073 | 891 | 913 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.444 | 892 | 921 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0021 | 893 | 912 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.004 | 914 | 936 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.542 | 915 | 939 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.7E-4 | 917 | 936 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1090 aa Download sequence Send to blast |
MADARRYGLT PQLGTILSRY FWKHNIDGYA LQKFVKFFET IRNFALPQNR QINLLDLIIS 60 FVLRGRSAGG SLFLFDRKVL RYFRKDGHNW RKKKDGKTVK EAHERLKAGS IDVLHCYYAH 120 GEENENFQRR SYWMLEEDYM HIVLVHYLEV KGNKPSFSRA RDVDEIAQVA NMDSPVCSNS 180 FTNHSQLPSQ TTSAESPNST HTSEYEDAES DNYQASSRYN SFLEMQQHGD GPVMDVRLLN 240 PHVPIDSVNN QCENDSDIQG TKATEPKSDF YSVLQENITR VFDETGLGLT FSGPRTQFDL 300 TSWDEVLEHC TTGFQAPSFY PAVASTQSAT VEDNLRLETS TLGELHTDDL GFKQVDVTSA 360 QDKSLWQLSN ADIGPLVTPN VDLQHGTSIE ENVNAPSLIT QASLDFSNIE GEGLKKYDSF 420 SRWMSNELGE VDDSHMKPSS GLYWNTVESE SVVEDSSMSN REHFDAYIMN PSLSQDQLFS 480 IIDFTPNWAY AGMETKVLIT GTFLKNKEDV EKCQWSCMFG EIEVPAEILT DGTLRCHAPL 540 HKSGRVPFYV TCSNRLACSE VREFEFREND AQYMEASDSY GYNTNEMCLH IRLEKLLTLG 600 PVDHQKAVPD SAKENLHLRN KISSLMMEAN DEWSNLVKLT HEGFSPDNAK DQLLEKLMKE 660 KLHSWLLHKV SEGGKGPNVL DKEGQGVLHL AAALGYDWAI RPTITAGVSI NFRDVHGWTA 720 LHWAANYGRE RTVVALIALD AAPGALTDPT PEYPTGRTPA DLASANGHKG IAGFLAESSL 780 TNHLSTLTLK ESKGSDVAEI SGITDVEDVA EKSAIQVADG DVQAGLSLKD SLSAVRNASL 840 AAARIYQVFR VHSFHRKKVI EYGDDKCGIS DERALSLISL KTAKPGQHDM PLHAAAIRIQ 900 NKFRGWKGRK EFLIIRQRIV KIQAHVRGYQ VRKQYKKIVW SVLIVEKAIL RWRRKGSGLR 960 GFRPEGQLEG PTMQNQGAKE DDYDFLQEGR KQTEARLQKA LARVKSMVQY PEARDQYRRL 1020 LNVVTELQES KKYDENTTRH LCVEGCQLVD QFVGKAMQDR ILKESEEAAA DDFMIELEEL 1080 WQDDTLMPTA |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
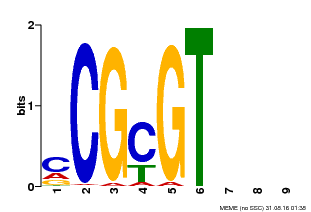
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008795548.1 | 0.0 | calmodulin-binding transcription activator 3-like isoform X2 | ||||
| TrEMBL | A0A2H3Y9P0 | 0.0 | A0A2H3Y9P0_PHODC; calmodulin-binding transcription activator 3-like isoform X2 | ||||
| STRING | XP_008795547.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



