 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s1158611g004 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 249aa MW: 28391.4 Da PI: 5.1287 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.6 | 2.2e-18 | 32 | 86 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ +q+++Le+ Fe ++++ +++ +LA++lgL+ rqV +WFqNrRa++k
PDK_30s1158611g004 32 EKKRRLSVYQVKALEKNFEVENKLEPDRKVRLARELGLQPRQVAIWFQNRRARWK 86
677899************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 127.8 | 4.4e-41 | 32 | 123 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
ekkrrls qvk+LE++Fe e+kLep+rKv+lareLglqprqva+WFqnrRAR+ktkqlE+dy+aLk +y+al+ + ++L++++e+L +e+
PDK_30s1158611g004 32 EKKRRLSVYQVKALEKNFEVENKLEPDRKVRLARELGLQPRQVAIWFQNRRARWKTKQLERDYAALKGSYEALRLDFHSLHRDKEALLAEM 122
69*************************************************************************************9998 PP
HD-ZIP_I/II 92 k 92
k
PDK_30s1158611g004 123 K 123
7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.07E-20 | 19 | 90 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.7E-20 | 25 | 88 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.896 | 28 | 88 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.1E-15 | 31 | 86 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 4.9E-17 | 31 | 92 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.17E-17 | 31 | 89 | No hit | No description |
| PRINTS | PR00031 | 9.4E-6 | 59 | 68 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 63 | 86 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.4E-6 | 68 | 84 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.1E-14 | 88 | 129 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 249 aa Download sequence Send to blast |
MYERGFQPII DGLDEEECGD GEMCSSAGSG REKKRRLSVY QVKALEKNFE VENKLEPDRK 60 VRLARELGLQ PRQVAIWFQN RRARWKTKQL ERDYAALKGS YEALRLDFHS LHRDKEALLA 120 EMKMLKDKLA EEESASFSSV KEEPLTGRSS DSDSSAVLND ENSPHDHRRM SSSTASGMFS 180 AAIGFENSTS FPCSPPSLLN SDSRQQKGFQ HQLLKTEDDE FLSGEEPCSS FFSDDHAATL 240 SWYYSEHWT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 80 | 88 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
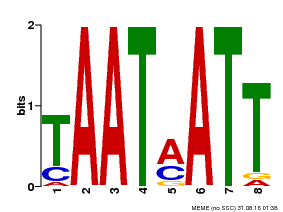
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated in leaves by drought stress. {ECO:0000269|PubMed:17999151}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT036856 | 2e-37 | BT036856.2 Zea mays full-length cDNA clone ZM_BFb0143M01 mRNA, complete cds. | |||
| GenBank | BT054500 | 2e-37 | BT054500.1 Zea mays full-length cDNA clone ZM_BFc0103D23 mRNA, complete cds. | |||
| GenBank | JX469932 | 2e-37 | JX469932.1 Zea mays subsp. mays clone UT3191 HB-type transcription factor mRNA, partial cds. | |||
| GenBank | JX514832 | 2e-37 | JX514832.1 Zea mays cultivar B73 homeodomain leucine zipper protein 10 (hdz10) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_026660788.1 | 1e-174 | homeobox-leucine zipper protein HOX20-like isoform X2 | ||||
| Swissprot | A2YWC0 | 5e-64 | HOX20_ORYSI; Homeobox-leucine zipper protein HOX20 | ||||
| Swissprot | Q6Z248 | 5e-64 | HOX20_ORYSJ; Homeobox-leucine zipper protein HOX20 | ||||
| TrEMBL | A0A3Q0I1M1 | 1e-173 | A0A3Q0I1M1_PHODC; homeobox-leucine zipper protein HOX20-like isoform X2 | ||||
| STRING | XP_008790926.1 | 1e-174 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3009 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G40060.1 | 4e-58 | homeobox protein 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



