 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s1016151g001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 295aa MW: 33427.6 Da PI: 5.811 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 165.5 | 1.8e-51 | 21 | 148 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkev 92
pGfrFhPtdeelv +yL++kve+k+l++ ++ike+diyk++PwdLp++++++ekewyfF+ r +ky+++ r+nr+t sg+Wkatg d+++
PDK_30s1016151g001 21 LPGFRFHPTDEELVGFYLRRKVEKKPLSI-DMIKEIDIYKYDPWDLPTSTAGGEKEWYFFCLRGRKYRNSIRPNRVTGSGFWKATGIDRPI 110
69***************************.99***************8888999************************************* PP
NAM 93 lsk..kgelvglkktLvfykgrapkgektdWvmheyrl 128
+s+ ge +glkk+Lv+y+g+a kg+ktdW+mhe+rl
PDK_30s1016151g001 111 YSAgdAGECIGLKKSLVYYRGSAGKGTKTDWMMHEFRL 148
**9777788***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 3.79E-56 | 19 | 175 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 55.03 | 20 | 175 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.7E-25 | 22 | 148 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005992 | Biological Process | trehalose biosynthetic process | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006561 | Biological Process | proline biosynthetic process | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0010120 | Biological Process | camalexin biosynthetic process | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MDSKSAGGEQ EVVREEEDVV LPGFRFHPTD EELVGFYLRR KVEKKPLSID MIKEIDIYKY 60 DPWDLPTSTA GGEKEWYFFC LRGRKYRNSI RPNRVTGSGF WKATGIDRPI YSAGDAGECI 120 GLKKSLVYYR GSAGKGTKTD WMMHEFRLPA SGKNDDISPS AQEAEIWTIC RIFKRNISCR 180 KYPQDMKAST SKQMPTDSGS KTSSFESDTG DEYKCFGSSS IPQGEMKLIP SYYEEKNQLL 240 AGQWNSAAQA PLATLHPNIP NMSTIEFFKD SNWDELGRIM EFMPDPSAAH ECGYT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 2e-50 | 5 | 181 | 2 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 2e-50 | 5 | 181 | 2 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 2e-50 | 5 | 181 | 2 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 2e-50 | 5 | 181 | 2 | 171 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 2e-50 | 5 | 181 | 2 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 2e-50 | 5 | 181 | 2 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
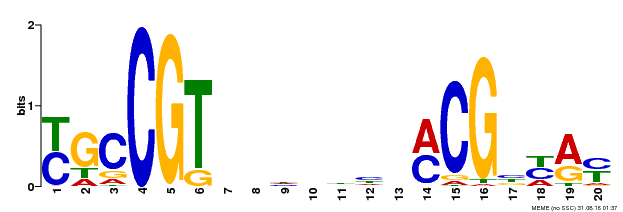
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00310 | DAP | Transfer from AT2G43000 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008806688.1 | 0.0 | transcription factor JUNGBRUNNEN 1-like isoform X1 | ||||
| Swissprot | Q9SK55 | 3e-90 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | A0A2H3Z0S9 | 0.0 | A0A2H3Z0S9_PHODC; transcription factor JUNGBRUNNEN 1-like isoform X1 | ||||
| STRING | XP_008806688.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1191 | 38 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43000.1 | 3e-91 | NAC domain containing protein 42 | ||||



