 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr008557.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 362aa MW: 40942.2 Da PI: 7.3902 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 29.6 | 1.5e-09 | 75 | 117 | 3 | 45 |
XXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 3 elkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
++k+ rr+++NReAAr+sR RKka++++Le+ +L + ++L
Pbr008557.1 75 ADKARRRLEQNREAARKSRIRKKAYVQQLETSRLKLAKLEQEL 117
7899**************************9666666555544 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 6.2E-9 | 71 | 122 | No hit | No description |
| SMART | SM00338 | 4.2E-5 | 73 | 144 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.14 | 75 | 119 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 1.4E-6 | 75 | 118 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 5.71E-7 | 77 | 115 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 80 | 95 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 4.9E-27 | 166 | 238 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 362 aa Download sequence Send to blast |
MSYPSTQFAT SRQMGIYEPF RQVDVWKETL RGDHGSNTAA SSIIEVDSGI ETKFGFVSHE 60 SMMPSGNDHE PNRSADKARR RLEQNREAAR KSRIRKKAYV QQLETSRLKL AKLEQELDKA 120 RKQGAYAGNA LGTSHMGYNG ALNSGITTFE MEYGHWVEEQ YRQNVELRNA LQDHVTDIEL 180 LKLVKGALNH YANLFRMKAD AATADVFYLL SGIWRTSVER YFHWIGGFRP SELLNIVLPQ 240 LQPLTEQQLL DFCNLRLSSQ QAEDALSQGM DKLQQTLALS IAADQMSGES FGSQMASALE 300 KLEALENFVS QADHLRQQTL QQMSRILTTY QAAQALLAMG EYFHRLRALS SLWTARPREP 360 T* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
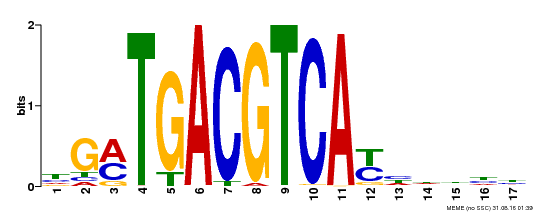
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. May be involved in the induction of the systemic acquired resistance (SAR) via its interaction with NPR1 (By similarity). {ECO:0000250, ECO:0000269|PubMed:12953119}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00247 | DAP | Transfer from AT1G77920 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr008557.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009367344.1 | 0.0 | PREDICTED: transcription factor TGA7 isoform X3 | ||||
| Refseq | XP_018505330.1 | 0.0 | PREDICTED: transcription factor TGA7 isoform X4 | ||||
| Swissprot | Q93ZE2 | 1e-155 | TGA7_ARATH; Transcription factor TGA7 | ||||
| TrEMBL | A0A498J846 | 0.0 | A0A498J846_MALDO; Uncharacterized protein | ||||
| STRING | XP_009367342.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1096 | 34 | 104 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G77920.1 | 1e-143 | bZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



