 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf01071g00011.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1034aa MW: 115422 Da PI: 6.0403 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 106.5 | 1.8e-33 | 20 | 95 | 2 | 77 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvgg 77
l+ ++rwl++ ei++iL n++ ++++ e+++rp+sgsl+L++rk++ryfrkDG+sw+kkkdgktv+E+he+LK +
Peaxi162Scf01071g00011.1 20 LEAQHRWLRPAEICEILRNYQTFKIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHSWRKKKDGKTVKEAHERLKGNR 95
5679*********************************************************************765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 48.573 | 15 | 159 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.3E-37 | 18 | 123 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.5E-28 | 21 | 96 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 7.0E-5 | 445 | 530 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 6.53E-16 | 446 | 532 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 9.3E-5 | 446 | 524 | IPR002909 | IPT domain |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-19 | 630 | 744 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.678 | 632 | 743 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.73E-19 | 633 | 743 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.60E-14 | 637 | 740 | No hit | No description |
| Pfam | PF12796 | 9.8E-9 | 654 | 744 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.0011 | 682 | 711 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.392 | 682 | 714 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 120 | 721 | 750 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.39E-6 | 853 | 908 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 23 | 857 | 879 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.492 | 859 | 887 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.14 | 860 | 878 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0042 | 880 | 902 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.285 | 881 | 905 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.6E-4 | 883 | 902 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1034 aa Download sequence Send to blast |
MANSRGYALN AQLDINQILL EAQHRWLRPA EICEILRNYQ TFKIAPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHSWRKKKD GKTVKEAHER LKGNRTNFSR IREPQQATPD FQETDDVHSS 120 EVESSASAKF YPNDYQVNSQ VTDTTSLSSA QASEYEDAES VYNQHSTSGF HSFLDAQPSM 180 MQNGGDAVPY RPVPFSNDHQ VQFAGSSDMN FSSIAPGNKG GNIENTYVPS RNLDLSWENI 240 SVNNPAAYQS YLYQPSSQSD ANNLMHEQGN ATLGQVFSND LKREEPQNRT DGLGDWRTSE 300 GGSSFISKWS MDQKLNPDLA SGGQQHKHPM QNELQSQLPD ANVGGSLSAD LHHNLSVGIK 360 ADYPAIKQPL LDGVLRAGLK KLDSFDRWMS KELEDVSEPH MQSNSSSYWD NVGDEEGVDN 420 STIASQVQLD SYMLSPSLSQ DQFFSIIDFS PNWAFAESEI KVLITGKFLK SQQEVEKCSW 480 ACMFGELEVP AEVIADGVLR CHTPIQKAGR VPFYITCSNR LACSEVREFE FRVTDVQDVD 540 VTNANTCSSS ESLLHMRFGK LLSLESTASQ NSPPSCEDDV SHMCSNINSL LKEDDSEWDE 600 MLHLTYENNF MAEKVKDQLL QKLLKEKLRV WLLQKVVEGG KGPNVLDEGG QGVLHFAAAL 660 GYDWAIPPTI AAGVSVNFRD VNGWTALHWA ASYGRLRTVG FLISLGAAPG ALTDPTPKHP 720 SGRTPADLAS SNGHKGIAGY LAESSLSSHL SFLGLKEKNQ VENVQAFGVE DVQTASERTA 780 TPAWEGDWPH GVSLKDSLAA VRNATQAAAR IHQVFRVQSF QRKQLKEYGG SEFGLSDERA 840 LSLLALKTKR AGHHDEPVQA AAVRIQNKFR SWKGRRDYLL IRQRIIKIQA HVRGHQVRNK 900 YKNIIWSVGI LEKVILRWRR KGSGLRGFKP EALAEGSNTQ DQPVQEDDYD FLKEGRKQTE 960 VRLQKALARV KSMVQYPEAR DQYRRLLNVV SDMKEPKAAN DGAPSRSVEA ADFGDDLIDL 1020 DDLLDDDTFM PTAP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4hb5_A | 3e-12 | 633 | 741 | 20 | 122 | Engineered Protein |
| 4hb5_B | 3e-12 | 633 | 741 | 20 | 122 | Engineered Protein |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
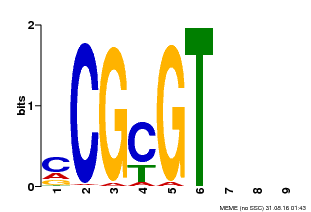
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013481 | 0.0 | BT013481.1 Lycopersicon esculentum clone 132164R, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009613616.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3 isoform X2 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A1S3YKL0 | 0.0 | A0A1S3YKL0_TOBAC; Calmodulin-binding transcription activator 4 | ||||
| STRING | XP_009613615.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2230 | 21 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



