 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf01009g00222.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 407aa MW: 46629 Da PI: 5.0394 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 113.1 | 1.8e-35 | 14 | 105 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksF 86
F+ k+ye+++d++++ ++sws++++sf+v+++ +fa+++LpkyFkh+nf+SF+RQLn+YgF+k++ e+ weF+++ F
Peaxi162Scf01009g00222.1 14 FIAKIYEMVDDPSTDAIVSWSSSNRSFIVWNPPDFARELLPKYFKHNNFSSFIRQLNTYGFRKTDPEQ---------WEFANEDF 89
9**************************************************************99988.........******** PP
XXXXXXXXXXXXXXXX CS
HSF_DNA-bind 87 kkgkkellekikrkks 102
+++ +ll++i+r+k
Peaxi162Scf01009g00222.1 90 LRDQPHLLKNIHRRKP 105
*************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 4.3E-39 | 5 | 98 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 6.39E-36 | 10 | 103 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 1.6E-60 | 10 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.8E-20 | 14 | 37 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 6.6E-32 | 14 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.8E-20 | 52 | 64 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.8E-20 | 65 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 407 aa Download sequence Send to blast |
MDEAACSTNA LPPFIAKIYE MVDDPSTDAI VSWSSSNRSF IVWNPPDFAR ELLPKYFKHN 60 NFSSFIRQLN TYGFRKTDPE QWEFANEDFL RDQPHLLKNI HRRKPVHSHS TQNLHSLPST 120 LNESERQAFK EDIENLKHEN ELLHLDLRRR KQDHQGLELK MQVFTERVQH VKHRQKNMLS 180 TLAQTISKPG LALSLMPQLE MNERKRRLPG NSYLYSETGL EDNQVCTSQV MTRENKDPTS 240 LLTFNKELLD QLEYSLTFWE NVLHDTDQSG MRRSSSVDLD ESISCADSPA ISYPQLNVDI 300 GSKFSAIDMN SEPNGNTIPE VAPPDDQVAV VGTATNVPTG VNDVFWEQFL TENPGSTDAP 360 EVKPERRDTD CEEAESQTVE HGKFWWKMKT VNSLAEQLGH LTPAEKT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 6e-26 | 4 | 103 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 6e-26 | 4 | 103 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 6e-26 | 4 | 103 | 19 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 202 | 207 | ERKRRL |
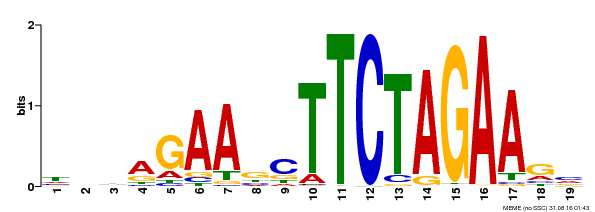
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019253075.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| Refseq | XP_019253077.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| Refseq | XP_019253078.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| Refseq | XP_019253079.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| Refseq | XP_019253080.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| TrEMBL | A0A1J6IT18 | 0.0 | A0A1J6IT18_NICAT; Heat stress transcription factor a-4a | ||||
| STRING | XP_009590974.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3448 | 23 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 1e-111 | heat shock transcription factor A4A | ||||



