 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MA_19683g0010 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Picea
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 341aa MW: 37932.8 Da PI: 6.7947 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 55.8 | 1.1e-17 | 46 | 95 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+WvAeIr p gk++r +lg+f tae+Aa+a+++a+ l+g
MA_19683g0010 46 QYRGVRQRS-WGKWVAEIRQP---GKQTRRWLGTFATAEQAAQAYDNAAILLYG 95
59****999.**********8...556999******************987766 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 2.6E-29 | 46 | 103 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.54E-29 | 46 | 105 | No hit | No description |
| PROSITE profile | PS51032 | 21.601 | 46 | 103 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.05E-19 | 46 | 103 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 1.1E-34 | 46 | 109 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 6.8E-10 | 47 | 58 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.0E-11 | 47 | 93 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 6.8E-10 | 69 | 85 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006970 | Biological Process | response to osmotic stress | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0009749 | Biological Process | response to glucose | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010353 | Biological Process | response to trehalose | ||||
| GO:0010449 | Biological Process | root meristem growth | ||||
| GO:0010896 | Biological Process | regulation of triglyceride catabolic process | ||||
| GO:0031930 | Biological Process | mitochondria-nucleus signaling pathway | ||||
| GO:0032880 | Biological Process | regulation of protein localization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 341 aa Download sequence Send to blast |
MKGGNREARC SAVGQENGGL RSTPSAPPSR KRICKGKGGP DNIKFQYRGV RQRSWGKWVA 60 EIRQPGKQTR RWLGTFATAE QAAQAYDNAA ILLYGSRAHL NLQPSGWDHS KSSSHTSKLR 120 PLLPRFTFRL PPAIHGTNPN PNPNPNCFGA IPSGYFQTPT NPDFWPAAMR AAQTDATYNL 180 PVIDPKRETT LEQSSVLHLE EVRVPTQQCN AEVHYESSSL KIDNLQENYG AEITDLQSHK 240 LGGPDQLSGI HGGVVGDELH QGFYSDTTGN IIEPRNHIDG ESRDDNLAAS LQELQYSAAP 300 PSPGFMWHYN IKHDYYYEET NSSTLPETNN QLWDYSDESS I |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 5e-16 | 46 | 102 | 2 | 59 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00674 | SELEX | Transfer from GRMZM2G093595 | Download |
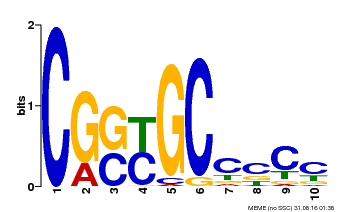
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | G0ZAE4 | 0.0 | G0ZAE4_PINTA; Putative ABI4-like protein | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40220.1 | 2e-25 | ERF family protein | ||||



