 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_15375A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 387aa MW: 43680.9 Da PI: 8.7392 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 22.6 | 2.8e-07 | 88 | 110 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C++Cg +F+ + +Lk+H+++H
Oropetium_20150105_15375A 88 HVCEKCGIRFKKPAHLKQHMQSH 110
79*******************99 PP
| |||||||
| 2 | zf-C2H2 | 17.4 | 1.2e-05 | 116 | 140 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C+ C+ +srk++L+rH+ tH
Oropetium_20150105_15375A 116 FACHvdGCPLNYSRKDHLNRHLLTH 140
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 14.5 | 0.0001 | 145 | 170 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp C++ Fs k n +rH H
Oropetium_20150105_15375A 145 FSCPieGCSRKFSVKGNVQRHVEEiH 170
89*******************98777 PP
| |||||||
| 4 | zf-C2H2 | 20.4 | 1.4e-06 | 181 | 205 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp +Cgk+F+ s Lk+H +H
Oropetium_20150105_15375A 181 FICPevNCGKVFKYASKLKKHEESH 205
78*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 15.8 | 4e-05 | 272 | 296 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ dC+ sFs ksnL++H + H
Oropetium_20150105_15375A 272 KCHfkDCKCSFSKKSNLNKHVKAaH 296
7*******************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 15 | 43 | 65 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.36 | 88 | 115 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.1E-5 | 88 | 106 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0021 | 88 | 110 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 90 | 110 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.94E-11 | 101 | 142 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-13 | 107 | 142 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.031 | 116 | 140 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.078 | 116 | 145 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 118 | 140 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.26E-6 | 127 | 177 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 4.3E-5 | 143 | 167 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0032 | 145 | 170 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.783 | 145 | 175 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 147 | 170 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 9.67E-5 | 177 | 207 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.8E-6 | 177 | 205 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.053 | 181 | 205 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0036 | 181 | 205 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 183 | 205 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 4.9 | 213 | 238 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 215 | 238 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 8.9 | 241 | 262 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-4 | 252 | 293 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.94E-6 | 252 | 313 | No hit | No description |
| PROSITE profile | PS50157 | 11.406 | 271 | 301 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.12 | 271 | 296 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 273 | 296 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-5 | 295 | 324 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 7 | 302 | 328 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.85 | 302 | 333 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 304 | 328 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 387 aa Download sequence Send to blast |
MGNGEPVTGG GEEAGGCEEA NTGAAASGAA AATPAPVRDI RRYKCEFCSV VRSKKSLIRA 60 HVLEHHKDEV DGLKDYQEGG VPRKEVFHVC EKCGIRFKKP AHLKQHMQSH SLERPFACHV 120 DGCPLNYSRK DHLNRHLLTH QGKLFSCPIE GCSRKFSVKG NVQRHVEEIH KDGSPCEKKE 180 FICPEVNCGK VFKYASKLKK HEESHVKLDY MEVICCEPGC MKTFTNVECL VAHKQSCHRH 240 VQCDVCGAKQ LKKNFKRHQR MHEDSCVTEK IKCHFKDCKC SFSKKSNLNK HVKAAHEQVK 300 PFLCQFSGCG KKFSYKHVRD NHEKSSAHVY TEGDFEEADE QRRHSAGGRK RNRITIESLM 360 RKRIAASDDA PAPEDGIEYL RWLLSG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 3e-16 | 93 | 225 | 20 | 147 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 3e-16 | 93 | 225 | 20 | 147 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
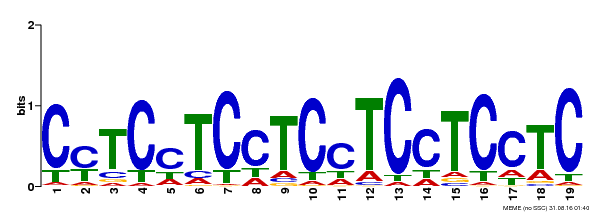
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_15375A |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004960333.1 | 0.0 | transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 1e-115 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | K3Z744 | 0.0 | K3Z744_SETIT; Uncharacterized protein | ||||
| STRING | Si022364m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-110 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_15375A |



