 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_11651A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 701aa MW: 77335 Da PI: 6.6581 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.8 | 4.7e-24 | 140 | 241 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegD 78
f+k+lt sd++++g +++p++ ae++ +++ + s++l+ +d++g +W++++iyr++++r++lt+GW+ Fv+ ++L +gD
Oropetium_20150105_11651A 140 FCKTLTASDTSTHGGFSVPRRAAEDCfppleySLQRP-SQELVAKDLHGTEWKFRHIYRGQPRRHLLTTGWSAFVNKKKLVSGD 222
99****************************9866654.569******************************************* PP
EEEEEE-SSSEE..EEEEE-S CS
B3 79 fvvFkldgrsefelvvkvfrk 99
v+F + +++ l+++v+r+
Oropetium_20150105_11651A 223 AVLFL--RGEDGVLRLGVRRA 241
*****..34899899999986 PP
| |||||||
| 2 | Auxin_resp | 99.2 | 5.7e-33 | 267 | 347 | 1 | 82 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsL 82
+a+a+++k+vF+++YnPr s+seF+++++k+++++++++s+G++f++ +e+ed++err++G+++g +++ p W++SkW++L
Oropetium_20150105_11651A 267 VARAVAMKTVFHICYNPRLSQSEFIIPYWKFMRSFNQPFSPGTKFRLVYESEDAPERRCTGVIIGSREAGP-MWHDSKWKCL 347
68899***************************************************************999.9********9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 6.3E-39 | 136 | 243 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.06E-40 | 136 | 246 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 8.58E-23 | 138 | 240 | No hit | No description |
| PROSITE profile | PS50863 | 13.338 | 140 | 242 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 6.8E-22 | 140 | 241 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 8.1E-23 | 140 | 242 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 8.3E-27 | 268 | 347 | IPR010525 | Auxin response factor |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009725 | Biological Process | response to hormone | ||||
| GO:0010050 | Biological Process | vegetative phase change | ||||
| GO:0010158 | Biological Process | abaxial cell fate specification | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 701 aa Download sequence Send to blast |
MTGIDLNTVE EEDEEGEAPQ APVAPGRADG AVCLELWHAC AGPVPPLPRK GSAVVYLPQG 60 HLEHISGGGD EAPAGVAAVP PHVFCRVVDV DLHADAATDE VYARVSLAPD DEAAAREYGS 120 AADRDDDAVT KRLAPVPHMF CKTLTASDTS THGGFSVPRR AAEDCFPPLE YSLQRPSQEL 180 VAKDLHGTEW KFRHIYRGQP RRHLLTTGWS AFVNKKKLVS GDAVLFLRGE DGVLRLGVRR 240 AAQLKDIHPT TPALHNQCSG NRNLENVARA VAMKTVFHIC YNPRLSQSEF IIPYWKFMRS 300 FNQPFSPGTK FRLVYESEDA PERRCTGVII GSREAGPMWH DSKWKCLVVR WDDNIEYRRP 360 NRVSPWEIEL TRSVSGSHLS PNPKRLKPCL PQVKAGTMIP DGSISSDFVG STRFHKVLQG 420 QELLGFKTHD GTAISASQAI EARNFRYNNE HRSSVNMSDI LGIPRHFVRS PYGIPGFSYH 480 CFGFGESQRF QKVLQGQEVL HPFRGGLGDA RIRTAGMYQS DGTHASGAGY KWLAPQGCDF 540 QQPAQPVVLQ ASSPSSVLMF PQASSKMPHF EHEYSCLDKD ATTQNLARTD QALSLWPRLI 600 SGEVTDEYTG AEKLHSPVSD AEHESNNEST VENGCRIFGI SLAEKVRSCD EADSCNVKYN 660 SLLLGSNLQM PKPLGTCWAT GLWQTSFQDI EVDVIVIHET * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-105 | 26 | 377 | 13 | 361 | Auxin response factor 1 |
| 4ldw_A | 1e-105 | 26 | 377 | 13 | 361 | Auxin response factor 1 |
| 4ldw_B | 1e-105 | 26 | 377 | 13 | 361 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00021 | PBM | Transfer from AT2G33860 | Download |
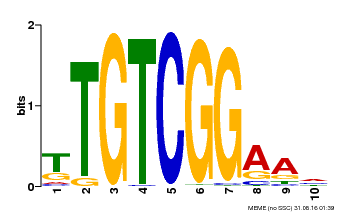
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_11651A |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin under dark condition. {ECO:0000269|PubMed:17408882}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025818787.1 | 0.0 | auxin response factor 2-like | ||||
| Swissprot | Q0JKI9 | 0.0 | ARFB_ORYSJ; Auxin response factor 2 | ||||
| TrEMBL | A0A2T7DIM0 | 0.0 | A0A2T7DIM0_9POAL; Auxin response factor | ||||
| STRING | Pavir.Eb02716.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2537 | 35 | 80 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33860.1 | 1e-143 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_11651A |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



