 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_07708A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 331aa MW: 36748.7 Da PI: 7.2049 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.7 | 2.2e-09 | 227 | 282 | 3 | 55 |
--SS--HHHHHHHHHHHH...HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFe...knrypsaeereeLAkklgLterqVkvWFqNrRake 55
kR ++ ke + L e k +yps++e+ LA+ +gL+++q+ +WF N+R ++
Oropetium_20150105_07708A 227 KRGKLPKEARQKLLHWWElhhKWPYPSETEKIALAEATGLDQKQINNWFINQRKRH 282
555555555555554444332789*****************************885 PP
| |||||||
| 2 | ELK | 42.9 | 1e-14 | 202 | 223 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELKhqLlrKY+g+Lg+L+qEFs
Oropetium_20150105_07708A 202 ELKHQLLRKYGGCLGGLRQEFS 223
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 9.3E-20 | 75 | 119 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 3.2E-21 | 77 | 116 | IPR005540 | KNOX1 |
| SMART | SM01256 | 3.4E-27 | 121 | 172 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 3.2E-23 | 125 | 170 | IPR005541 | KNOX2 |
| SMART | SM01188 | 4.9E-8 | 202 | 223 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.034 | 202 | 222 | IPR005539 | ELK domain |
| Pfam | PF03789 | 3.6E-11 | 202 | 223 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 13.394 | 222 | 285 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.21E-20 | 223 | 295 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 3.2E-14 | 224 | 289 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.36E-15 | 225 | 285 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-28 | 227 | 288 | IPR009057 | Homeodomain-like |
| Pfam | PF05920 | 3.6E-16 | 242 | 281 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 260 | 283 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 331 aa Download sequence Send to blast |
MKRFAYLGEG GSSSTSNASF LQLPLPTAAS ASASAQVIPP EAHRSRLPLQ QLLADPSSSV 60 QTVRQSHQKG EISPPDIKAK IMSHPQYSAL VTAYLDCQKV GAPPDVSDRL SAVAVKLDQA 120 QRWPPRADPE LDQFMEAYCN MLVKYKEEMA RPIQEAAEFF KSVERQLDSI ADSSACDGAG 180 SSEDDQDTSC PEEVDPCAEV KELKHQLLRK YGGCLGGLRQ EFSKRKKRGK LPKEARQKLL 240 HWWELHHKWP YPSETEKIAL AEATGLDQKQ INNWFINQRK RHWKPTSEDM PFAMTTTMEG 300 GFILAPPPGA AAALYMDRPF MGDGMYRLGS * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during embryogenesis. {ECO:0000269|PubMed:10080693}. | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during embryogenesis. {ECO:0000269|PubMed:10080693, ECO:0000269|PubMed:9869405}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00671 | PBM | Transfer from GRMZM2G135447 | Download |
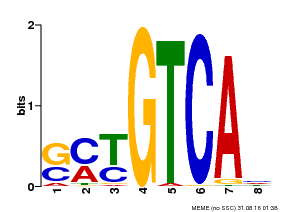
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_07708A |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025795146.1 | 1e-160 | homeobox protein knotted-1-like 12 isoform X3 | ||||
| Swissprot | O65034 | 1e-110 | KNOSC_ORYSI; Homeobox protein knotted-1-like 12 | ||||
| Swissprot | O80416 | 1e-110 | KNOSC_ORYSJ; Homeobox protein knotted-1-like 12 | ||||
| TrEMBL | A0A0A9GAR1 | 1e-173 | A0A0A9GAR1_ARUDO; Uncharacterized protein | ||||
| STRING | Sb01g006790.1 | 1e-151 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4061 | 38 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 7e-88 | KNOTTED-like from Arabidopsis thaliana | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_07708A |



