 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_01474A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 291aa MW: 32922.4 Da PI: 9.0833 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 99.5 | 1.3e-31 | 9 | 58 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
krienk+nrqvtfskRrng+lKKA+ELSvLCdaeva+i+fss+gklye+
Oropetium_20150105_01474A 9 KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFG 58
79**********************************************95 PP
| |||||||
| 2 | K-box | 15.1 | 9.3e-07 | 105 | 128 | 13 | 36 |
K-box 13 kaeslqqelakLkkeienLqreqR 36
+++s++qe+++Lk++ e+Lqr+qR
Oropetium_20150105_01474A 105 DSKSWYQEISRLKAKFESLQRSQR 128
5799******************99 PP
| |||||||
| 3 | K-box | 71 | 3.6e-24 | 149 | 213 | 35 | 99 |
K-box 35 qRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkkle 99
+Rh+lGedL++Ls+keLqqLe+qLe +l++ R++K+++++eq++el++ke++l e nk+L+++le
Oropetium_20150105_01474A 149 LRHMLGEDLGPLSIKELQQLEKQLEYALSQARQRKTQIMMEQMDELRRKERQLGELNKQLKNQLE 213
59************************************************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 33.529 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 1.6E-41 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 2.07E-45 | 2 | 78 | No hit | No description |
| SuperFamily | SSF55455 | 5.23E-33 | 2 | 76 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.0E-32 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 2.6E-27 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.0E-32 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.0E-32 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS51297 | 15.144 | 106 | 218 | IPR002487 | Transcription factor, K-box |
| Pfam | PF01486 | 7.4E-21 | 149 | 212 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048437 | Biological Process | floral organ development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 291 aa Download sequence Send to blast |
MGRGRVELKR IENKINRQVT FSKRRNGLLK KAYELSVLCD AEVALIVFSS RGKLYEFGSA 60 GVNKTLEKYH NCCYNHGQGS NNGFGGEPQL EVVRMHASKY VSLFDSKSWY QEISRLKAKF 120 ESLQRSQRTD DYSSRYLLHY VISTYFLFLR HMLGEDLGPL SIKELQQLEK QLEYALSQAR 180 QRKTQIMMEQ MDELRRKERQ LGELNKQLKN QLEAEGCSRY KAVETSCAPN ATVGIDGRAH 240 SVPNALPGAA TDCEPTLQIG YHQFAAPEPA AMPRNGTDGG ENNFVLGWVL * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3mu6_A | 3e-20 | 2 | 69 | 1 | 68 | Myocyte-specific enhancer factor 2A |
| 3mu6_B | 3e-20 | 2 | 69 | 1 | 68 | Myocyte-specific enhancer factor 2A |
| 3mu6_C | 3e-20 | 2 | 69 | 1 | 68 | Myocyte-specific enhancer factor 2A |
| 3mu6_D | 3e-20 | 2 | 69 | 1 | 68 | Myocyte-specific enhancer factor 2A |
| 5f28_A | 4e-20 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_B | 4e-20 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_C | 4e-20 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_D | 4e-20 | 1 | 69 | 1 | 69 | MEF2C |
| 6byy_A | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| 6byy_B | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| 6byy_C | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| 6byy_D | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| 6bz1_A | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| 6bz1_B | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| 6bz1_C | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| 6bz1_D | 4e-20 | 1 | 81 | 1 | 81 | MEF2 CHIMERA |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Plays minor but redundant roles with MADS6 in floral development. {ECO:0000269|PubMed:19820190}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00315 | DAP | Transfer from AT2G45650 | Download |
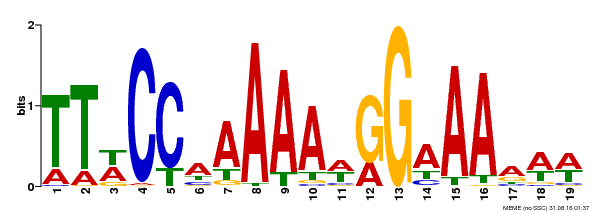
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_01474A |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY541067 | 9e-71 | AY541067.1 Hordeum vulgare subsp. vulgare AGAMOUS LIKE6-like protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025824169.1 | 1e-141 | MADS-box transcription factor 17-like | ||||
| Swissprot | Q7XUN2 | 1e-112 | MAD17_ORYSJ; MADS-box transcription factor 17 | ||||
| TrEMBL | A0A2S3I9M9 | 1e-139 | A0A2S3I9M9_9POAL; Uncharacterized protein | ||||
| STRING | Si010898m | 1e-137 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4187 | 35 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45650.1 | 7e-84 | AGAMOUS-like 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_01474A |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



