 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ote100006770191 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Ocimeae; Ocimum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1285aa MW: 143336 Da PI: 8.629 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.3 | 0.00026 | 1169 | 1194 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y C C++sFs+k +L+ H r+ +
Ote100006770191|100006770191 1169 YACDmeGCTMSFSSKQELTLHKRNiC 1194
789999****************9877 PP
| |||||||
| 2 | zf-C2H2 | 13.3 | 0.00025 | 1194 | 1216 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Ote100006770191|100006770191 1194 CPvkGCGKKFFSHKYLVQHRRVH 1216
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 12.2 | 0.00057 | 1252 | 1278 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
Ote100006770191|100006770191 1252 YVCTetGCGQTFRFVSDFSRHKRKtgH 1278
99********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 3.8E-15 | 22 | 63 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.035 | 23 | 64 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 6.7E-14 | 24 | 57 | IPR003349 | JmjN domain |
| SuperFamily | SSF51197 | 7.69E-26 | 109 | 129 | No hit | No description |
| SMART | SM00558 | 4.8E-50 | 170 | 339 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.331 | 173 | 339 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 7.69E-26 | 188 | 344 | No hit | No description |
| Pfam | PF02373 | 1.7E-37 | 203 | 322 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 4.1E-4 | 1164 | 1191 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 11 | 1169 | 1191 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.362 | 1192 | 1221 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1192 | 1216 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 1193 | 1220 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1194 | 1216 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.23E-9 | 1208 | 1250 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-8 | 1221 | 1246 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1222 | 1251 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1222 | 1246 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1224 | 1246 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.97E-8 | 1240 | 1274 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.4E-9 | 1247 | 1274 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.074 | 1252 | 1283 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.19 | 1252 | 1278 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1254 | 1278 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1285 aa Download sequence Send to blast |
MAAEVSGGGN IEVPPWLKSM AVAPEYHPTL AEFQDPIAYI FKIEKEASSY GICKIVPPVP 60 TAPRKTVVSN LNKSLLARSN ETGPTFTTRQ QQIGFCPRKH RPVQKPVWQS GERYTLAEFE 120 AKAKGFEKNY LKKYAKKALS SLEIETLYWN AMVDKPLSVD GGKKNEGGLT VGETEWNMRT 180 LSRENLSLLK FMKEEIPGVT SPMVYVAMMF SWFAWHVEDH DLHSMNYLHL GSGKTWYGVP 240 RAAAVAFEEV IREHGYGGEI NPLVTFATLG EKTTVMSPEV LLNAGVPCCR LVQNAGEFVV 300 TFPRAYHSGF SHGFNCGEAA NIATPEWLRV AKEAAIRRAA INCPPMVSHF QLLYDLALSL 360 CSRVPKSIAT EPRSSRLKDR KRGEGDKLIK ELFYQDMMQN NDMLNVLGKG SSIVLLPQNY 420 LNHSFSSNPM TGFQSTKSKL FPSLCSLGLE LKNACDNASD DFTLDKKQGL KQAKGSVAKR 480 TFSALFGIDI HSPAHAGETH CELKKASPCE QGLFSCVTCG ILCFACVAIV QPTEATAQYL 540 MSADCSTLSN LEESDNDLTH ICDPKASNEN LPSVFVLRSQ NGLSADQHID GGNVCSASDS 600 KAHKEPSSLG LLALAYGNFS DSEEEDSEAI VPAEGCGTSK SDDFRNEDAN VNTDPKADCT 660 TEISLLTSDS SAKFDPALAT SKDGEALYPE FSDGCNKSTS TLIKSNSLTC RFSRHQTEPQ 720 LDIPNSLSRK AVGTPITGLA PQEPSTFPFS SRLDEDSSRL HVFCLQHAMQ VRRRLSQFGG 780 ADVFLVCHPD YPKLESEAKK VAEELESNYI WSEVSFREAT DEDEEIIRRA LESENAIHGN 840 RDWAVKLGIN LFYSANLSRS PLYSKQMHYN FVVYNAFGRS SPTDSSASEG EREGGGLGRQ 900 KKITVAGKWC GKVWMSNQAH PLLLNKDSQQ HEQDFAVASW TKPDVKYVRR TERRPAAGLL 960 SAARKMSRKK KNDPESSSLM RGKSLEAEKL DESRDFSLSN NKIKRKRTSR MVKEENPESE 1020 NVVESSEESQ LSDSCKESES RRGIKRSRKE NCEAEEDSFE EERPLSKSWK QIKRKQRAKR 1080 REKETPDTAK NQKKSKKHEL NTLVEEEMEG GPSTRLRKRT TKPSTDSGAR LSKPKPVVKA 1140 QQRDAKTTKK SSASKSPSVG KIRDEEAEYA CDMEGCTMSF SSKQELTLHK RNICPVKGCG 1200 KKFFSHKYLV QHRRVHMEDR PLKCPWKGCK MTFKWAWART EHIRVHTGAR PYVCTETGCG 1260 QTFRFVSDFS RHKRKTGHAP KKARG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 6e-69 | 16 | 360 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 6e-69 | 16 | 360 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
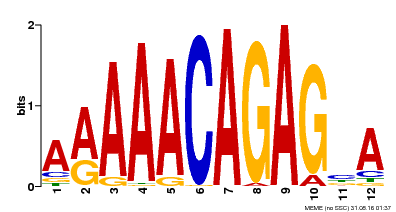
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011072922.1 | 0.0 | lysine-specific demethylase REF6 isoform X1 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A4D9ALJ0 | 0.0 | A0A4D9ALJ0_SALSN; Uncharacterized protein | ||||
| STRING | Migut.I00266.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4988 | 23 | 31 |



