 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os02g45670.1 | ||||||||
| Common Name | LOC9272080, Os02g0680700, OsJ_07933, OSNPB_020680700 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 292aa MW: 31834.9 Da PI: 9.2478 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 42.7 | 1.3e-13 | 37 | 81 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a +++ + Wk+I + +g +t q++s+ qky
LOC_Os02g45670.1 37 RESWTEQEHDKFLEALQLFDRD-WKKIEAFVG-SKTVIQIRSHAQKY 81
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.22E-17 | 31 | 87 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.408 | 32 | 86 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.3E-7 | 35 | 87 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.2E-18 | 35 | 84 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 5.3E-11 | 36 | 84 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.3E-11 | 37 | 81 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.52E-8 | 39 | 82 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0020039 | anatomy | leaf lamina | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 292 aa Download sequence Send to blast |
MVSANQPPPD AAAAAAGSAG EDASKKVRKP YTITKSRESW TEQEHDKFLE ALQLFDRDWK 60 KIEAFVGSKT VIQIRSHAQK YFLKVQKNGT SEHVPPPRPK RKAAHPYPQK ASKNEPGYTI 120 KADSSSMLRN SGMNATVSSW THNSIPPIVA SSMVKEDLGA GAMAPNNFCS SSTEGPARAW 180 QPGETNDQIN QVPSLRLMPD FAQVYSFLGS VFDPSTSGHL QKLKEMNPID VETALLLMRN 240 LSINLTSPDF EDQKKLLSSY STPSDGLELG STRSSVLADR PLSAPFMIKG E* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.12484 | 0.0 | callus| flower| leaf| panicle| seed| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 156764674 | 0.0 | ||||
| Expression Atlas | B9F1T8 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00625 | PBM | Transfer from PK02532.1 | Download |
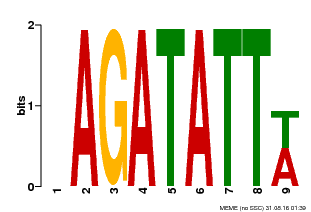
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os02g45670.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os02g45670 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK288059 | 0.0 | AK288059.1 Oryza sativa Japonica Group cDNA, clone: J075153L04, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015627665.1 | 0.0 | protein REVEILLE 6 isoform X2 | ||||
| Swissprot | Q8H0W3 | 1e-102 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | B9F1T8 | 0.0 | B9F1T8_ORYSJ; Os02g0680700 protein | ||||
| STRING | OS02T0680700-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1096 | 38 | 111 | Representative plant | OGRP928 | 15 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.1 | 9e-88 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os02g45670.1 |
| Entrez Gene | 9272080 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



