 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA007250-PA | ||||||||
| Common Name | OsI_05578 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 369aa MW: 42765.7 Da PI: 8.5261 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 18.8 | 4.5e-06 | 60 | 82 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cg sF + +Lk+H+++H
BGIOSGA007250-PA 60 HTCQECGASFQKPAHLKQHMQSH 82
79*******************99 PP
| |||||||
| 2 | zf-C2H2 | 22 | 4.3e-07 | 88 | 112 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s+ rk++L+rH+ +H
BGIOSGA007250-PA 88 FICPleDCPFSYIRKDHLNRHMLKH 112
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 15.6 | 4.6e-05 | 117 | 142 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C+ Cg+ Fs k n++rH + H
BGIOSGA007250-PA 117 FTCSmdGCGRKFSIKANMQRHVKEiH 142
89********************9988 PP
| |||||||
| 4 | zf-C2H2 | 15.8 | 3.9e-05 | 154 | 178 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ C+k+F+ s +k+H +H
BGIOSGA007250-PA 154 FVCKeeGCNKVFKYASKMKKHEESH 178
89*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 17.9 | 8.6e-06 | 245 | 269 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C sFs+ksnL++Hi+ H
BGIOSGA007250-PA 245 KCSfeGCECSFSNKSNLTKHIKAsH 269
688899***************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 24 | 14 | 36 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.9E-6 | 57 | 79 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 13.547 | 60 | 87 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0017 | 60 | 82 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 62 | 82 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.34E-11 | 72 | 114 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.1E-12 | 80 | 114 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 5.5E-4 | 88 | 112 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.808 | 88 | 117 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 90 | 112 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.55E-8 | 98 | 146 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-7 | 115 | 144 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 3.9E-4 | 117 | 142 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.154 | 117 | 147 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 119 | 142 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-7 | 151 | 178 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0038 | 154 | 178 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.764 | 154 | 178 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 156 | 178 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.7 | 186 | 211 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 188 | 211 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 214 | 235 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 8.2E-6 | 226 | 266 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 8.34E-8 | 227 | 282 | No hit | No description |
| SMART | SM00355 | 0.002 | 244 | 269 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.365 | 244 | 274 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 246 | 269 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.8 | 275 | 301 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 277 | 301 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 369 aa Download sequence Send to blast |
MRVEATQHRD IRRYKCEFCT VVRSKKCLIR AHMVAHHKEE LDKSEIYKSN GEKVVHEGDH 60 TCQECGASFQ KPAHLKQHMQ SHSDERSFIC PLEDCPFSYI RKDHLNRHML KHQGKLFTCS 120 MDGCGRKFSI KANMQRHVKE IHEDETATKS NRQFVCKEEG CNKVFKYASK MKKHEESHVK 180 LDYVEVVCCE PGCMKTFTNV ECLRAHNQAC HQYVQCDICG EKHLKKNIKR HLRAHEEVPS 240 TERIKCSFEG CECSFSNKSN LTKHIKASHD QVKPFACRFT GCEKVFPYKH VRDNHEKSSA 300 HVYTQANFTE MDEQLLSCPR GGRKRKAVTV ETLTRKRVTM HGDASSLDNG TEYLRWLLSG 360 GDDDSSQTH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 8e-19 | 64 | 250 | 19 | 174 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 8e-19 | 64 | 250 | 19 | 174 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.18086 | 0.0 | callus| flower| leaf| panicle| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32985682 | 0.0 | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Both isoforms 1 and 2 transcripts are expressed in seedlings and both TFIIIA and TFIIIA-C are present at similar ratios in 10-day-old seedlings (at protein level). In correlation with the reorganization of 5S rDNA chromatin to a mature state, full-length isoform 1 protein TFIIIA with transcriptional activity accumulates and permits de novo transcription of 5S rRNA. Isoforms 1 and 2 transcripts accumulate in various tissues of the reproductive phase, including flowers and siliques, but only isoform 2 is present in mature seeds. In flowers, both TFIIIA and TFIIIA-C are present at similar ratios (at protein level). Very low amounts of TFIIIA are found in extracts from fresh siliques compared to TFIIIA-C. The amount of full-length TFIIIA protein progressively increases from fresh siliques to seeds concomitant with lower proportions of the shorter TFIIIA-C protein (at protein level) thus leading to 5S rRNA accumulation in the seed. {ECO:0000269|PubMed:22353599}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in seedlings, flowers, siliques and seeds. {ECO:0000269|PubMed:22353599}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
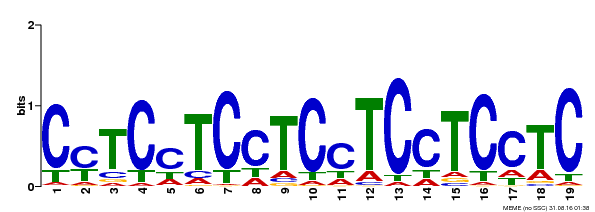
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK100473 | 0.0 | AK100473.1 Oryza sativa Japonica Group cDNA clone:J023093M20, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015626855.1 | 0.0 | transcription factor IIIA isoform X1 | ||||
| Swissprot | Q84MZ4 | 1e-105 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A0D3EZT1 | 0.0 | A0A0D3EZT1_9ORYZ; Uncharacterized protein | ||||
| TrEMBL | A2X037 | 0.0 | A2X037_ORYSI; Uncharacterized protein | ||||
| TrEMBL | A3A2F3 | 0.0 | A3A2F3_ORYSJ; Uncharacterized protein | ||||
| STRING | ORUFI02G01170.1 | 0.0 | (Oryza rufipogon) | ||||
| STRING | ORGLA02G0010700.1 | 0.0 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-108 | transcription factor IIIA | ||||



