 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA005011-PA | ||||||||
| Common Name | OsI_04849 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1286aa MW: 142117 Da PI: 8.8013 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.4 | 0.001 | 1192 | 1214 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L +H ++H
BGIOSGA005011-PA 1192 CPvkGCGKKFFSHKYLLQHRKVH 1214
9999*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 2.6E-15 | 24 | 65 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.201 | 25 | 66 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.5E-13 | 26 | 59 | IPR003349 | JmjN domain |
| SMART | SM00558 | 4.6E-51 | 198 | 367 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 37.486 | 201 | 367 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 4.94E-27 | 212 | 381 | No hit | No description |
| Pfam | PF02373 | 9.4E-39 | 231 | 350 | IPR003347 | JmjC domain |
| SMART | SM00355 | 17 | 1167 | 1189 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.612 | 1190 | 1219 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1190 | 1214 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 9.7E-6 | 1192 | 1218 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1192 | 1214 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.7E-9 | 1206 | 1246 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-8 | 1219 | 1244 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.323 | 1220 | 1249 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0027 | 1220 | 1244 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1222 | 1244 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.01E-7 | 1238 | 1272 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 8.5E-9 | 1245 | 1273 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.18 | 1250 | 1281 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.7 | 1250 | 1276 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1252 | 1276 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1286 aa Download sequence Send to blast |
MRPSPPPAAP AAEPVPPWLR SLPVAPEFRP TAAEFADPVS YILKIEPAAA PYGICKVVPP 60 LPPPPKKATF SNLSRSFAAL HPDDRSPSFP TRHQQVGLCP RRTRPGLKPV WRSSHRYTLP 120 QFESKAGATR KSLLAGLNFP ASRQLTPLDH EVLFWRASAD RPIVVEYGSD MSGSGFSPCA 180 AQPQPPPQQQ PTARAAAHLG ETAWNMRGVA RSPGSLLRFM PEDVPGVTTP MLYVGMMFSW 240 FAWHVEDHDL HSLNYMHLGA AKTWYGVPRD AALAFEDVVR EHGYGGEVNP LETFATLGQK 300 TTVMSPEVLV ESGIPCCRLV QNAGEFVVTF PGSYHCGFSH GFNCGEASNI ATPEWLRIAK 360 EAAIRRASIN RPPMVSHYQL LYDLALSMRF REPSNGEMET RSSRIKEKKK CEGEQLVKKM 420 FIQNVIEDNE LLSHLLNDGS SCIILPANAH DGPGLSTLRS TDQSNMNSRI SHNLCSREEA 480 PEASGCLSPN RNGDTRNCIS SDTHNMEGDK GDIMSATGLL DQGLLSCVTC GILSFSCVAV 540 LKPRDSTARY LMSADSNSIN NQFSISGGSI LADAPTNERN DVISRPYSEH CCNEIMADDA 600 EIDKNSALDL LAFAHGGQSD PEEDPLEKIL KIAHGINKSQ PNSSNNVGCV GTKLSSSSTE 660 RQERPSSQNA HCNGSSVISN GPKGVRTRNK YQLKMVLSEG FQAKDIYSAK EKKVQSEPSS 720 SKGDVKETID VSGTENDVGC KSTTISVSEH RGSTKNMYSV KENKVQSKPS SLKGTVKETV 780 DVSGTENDAR CKSITISVSE HRGSTPMTNS LAASIVKPDK DSSRMHVFCL EHAIEVEKQL 840 HAIGGSNIML ICRPEYPKIE AEARLLGEEM GLVYDWKGIH FKEANMEDRQ KIQEVLRDEE 900 AIPTSSDWAV KLGINLYYSA NLAKSPLYNK QMPYNRVIYR AFGCDSPNDS PVMFNTCERK 960 QSHQKKIVVA GRWCGKVWMS KQVHPYLAHR VESQEAEEAD RICSYHFDEK HKAEPVGNSS 1020 RVEASKRKSS SLTDVTESSN RRGEIPGEET NTKRPKHSQE NNLRALETAA EVVVPSPAGT 1080 GLRVSSRIAN RANKLKSKME KEDVPSSRPK SNIKEKSSHA SGQKSNVQEA NANSASHLRA 1140 MPPKQKAEAE AKKQIRTPKP PKQAVEYSCD IEGCSMSFRT KRDLSLHKSD ICPVKGCGKK 1200 FFSHKYLLQH RKVHTDDRPL TCPWKGCNMA FKWPWARTEH LRVHTGDRPY VCHEPGCAQT 1260 FRFVSDFSRH KRKTGHSVKK KKKAKS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 8e-74 | 18 | 381 | 8 | 346 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 8e-74 | 18 | 381 | 8 | 346 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.52727 | 0.0 | callus| panicle| root | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32978976 | 0.0 | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaves and flag leaves. Expressed at low levels in roots, shoots, stems and panicles. {ECO:0000269|PubMed:24280387}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3 with a specific activity for H3K27me3 and H3K27me2. No activity on H3K4me3, H3K9me3, H3K27me1 and H3K36me3. Involved in biotic stress response. May demethylate H3K27me3-marked defense-related genes and increase their basal and induced expression levels during pathogen infection. {ECO:0000269|PubMed:24280387}. | |||||
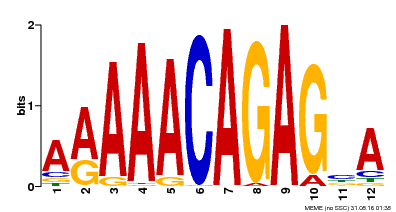
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt stress, abscisic acid (ABA), jasmonic acid (JA), the ethylene precursor ACC and infection by the bacterial pathogen Xanthomonas oryzae pv. oryzae. {ECO:0000269|PubMed:24280387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK068952 | 0.0 | AK068952.1 Oryza sativa Japonica Group cDNA clone:J023001N18, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015621377.1 | 0.0 | lysine-specific demethylase JMJ705 | ||||
| Swissprot | Q5N712 | 0.0 | JM705_ORYSJ; Lysine-specific demethylase JMJ705 | ||||
| TrEMBL | B8A7U6 | 0.0 | B8A7U6_ORYSI; Uncharacterized protein | ||||
| STRING | ONIVA01G46140.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5346 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



