 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI12G09490.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1366aa MW: 151845 Da PI: 7.4501 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.8 | 0.00035 | 1247 | 1270 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
++C C++sFs++ +L H r
ORUFI12G09490.1 1247 FPCDieGCDMSFSTQQDLLLHKRD 1270
789999***************985 PP
| |||||||
| 2 | zf-C2H2 | 11.9 | 0.0007 | 1330 | 1356 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
ORUFI12G09490.1 1330 YECQepGCGQTFRFVSDFSRHKRKtgH 1356
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.9E-16 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.201 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 3.4E-14 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 5.2E-43 | 183 | 352 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 32.998 | 183 | 352 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 9.89E-23 | 200 | 349 | No hit | No description |
| Pfam | PF02373 | 1.6E-34 | 216 | 335 | IPR003347 | JmjC domain |
| SMART | SM00355 | 9.7 | 1247 | 1269 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.907 | 1270 | 1299 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 1270 | 1294 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1272 | 1294 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.12E-14 | 1281 | 1335 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.3E-11 | 1297 | 1324 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 8.6E-4 | 1300 | 1324 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.655 | 1300 | 1329 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1302 | 1324 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.2E-10 | 1325 | 1353 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.926 | 1330 | 1361 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.18 | 1330 | 1356 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1332 | 1356 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1366 aa Download sequence Send to blast |
MSLQPPAVEP PEWLRTLPVA PEYHPTLAEF ADPIAYILRI EPEASRYGIC KIVPPLPRPP 60 EDDTFRRLQA AFAAAASSNG DPSPTFPTRL QQVGLSARNR RAASRRVWES GERYTLEAFR 120 AKAAEFEPPR HAAPPRNPTH LQLEALFWAA CASRPFSVEY GNDMPGSGFA SPDELPDAAN 180 ATDVGETEWN MRVAPRARGS LLRAMARDVA GVTTPMLYVA MLYSWFAWHV EDHELHSLNF 240 LHFGKAKTWY GVPRDAMLAF EETVRVHGYA DDLNAIMAFQ TLNEKTTVLS PEVLLSAGVP 300 CCRLVQKAGE FVITFPGAYH SGFSHGFNCG EASNIATPHW LQVAKEAAIR RASTNCGPMV 360 SHYQLLYELA LSLRPREPKN FYSVPRSSRL RDKNKNEGDI MVKENFVGSV TENNNLLSAL 420 LDKNSCIIVP NADFFVPSFP VALESEVTVK QRFTAGPCSI SQQGAENMAA DHVAVDKVTE 480 IQDMSGSLYP CETSLVGCSN RKLYETKYGQ RDAAALCLST SEIQSRGIDT ARSHPAGGIL 540 DQGRLPCVQC GILSFACVAI IQPREAAVQF IMSKECISSS AKQGGIGASD DTSNWIDQSH 600 EISPPPGPAS GTDDNVKHAV SLAHVSDRCR ELYASNTDGC TSALGLLASA YDSSDSDDET 660 TEDVSKHSKK NDSVNQSTDP QILETSASCS STVQCQKTNS HLHEEECEAR ATSLMKPVSH 720 NSRPISQSNR DTDIDHFIEL GKSGTQCSGY LDLVDDLTTS VLKSSSDTCV SAAKASMDPD 780 VLTMLRYNKD SCRMHVFCLE HALETWTQLQ QIGGANIMLL CHPEYPRAES AAKVIAEELG 840 IKHDWKDITF KEATEEDVKK IQLALQDEDA EPTGSDWAVK MGINIYYSAK QSKSPLYSKQ 900 IPYNSIIYKA FGQENPDSLT DYGCQKSGST KKKVAGWWCG KVWMSNQVHP LLAREREEQN 960 SSVVYGKAMF TTISHGKVQD EASTRCNTSN RTPSRRTSRR KKGVSAEKSK PKNKRSTASD 1020 EASMLCSGLG MNSGVIHDQT ENSDDYDKHG NGDEIEEGTN PQKYQQRKLQ NVTRKSSSKK 1080 RKDEKRTDSF HELYDEDNGV DYWLNMGSGD DATLGNSRQQ SPDPVKVKSG GKLQGKRKSS 1140 KYKSNDDLLN EENKLQKMNK KSSSKKQKND KINRQLQEDQ TEDDHMDHLV DVAVADEVTL 1200 DNEDKITEDK IDDVKVKSRG KSQNGKRKGS KHQATDGLRA GNKVAKFPCD IEGCDMSFST 1260 QQDLLLHKRD ICPVKGCKKK FFCHKYLLQH RKVHIDERPL KCTWKGCKKA FKWPWARTEH 1320 MRVHTGVRPY ECQEPGCGQT FRFVSDFSRH KRKTGHSSDK RRKNST |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 5e-70 | 12 | 373 | 7 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 5e-70 | 12 | 373 | 7 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
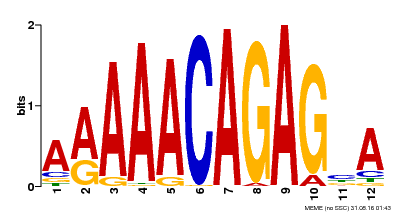
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK065251 | 0.0 | AK065251.1 Oryza sativa Japonica Group cDNA clone:J013002J08, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015620626.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A0E0RFZ5 | 0.0 | A0A0E0RFZ5_ORYRU; Uncharacterized protein | ||||
| TrEMBL | Q2QTX9 | 0.0 | Q2QTX9_ORYSJ; JmjC domain containing protein, expressed | ||||
| STRING | ORUFI12G09490.1 | 0.0 | (Oryza rufipogon) | ||||
| STRING | OS12T0279100-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6718 | 30 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-159 | relative of early flowering 6 | ||||



