 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI08G03840.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 747aa MW: 81729.1 Da PI: 5.9081 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 22.5 | 2.7e-07 | 24 | 50 | 1 | 28 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIa 28
r rWT+ E+ ++++a k++G W +I
ORUFI08G03840.1 24 RERWTEAEHNRFLEALKLYGRA-WQRIE 50
78******************88.**996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.2E-5 | 18 | 49 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 0.001 | 24 | 51 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 747 aa Download sequence Send to blast |
MEINSSGEEA VVKVRKPYTI TKQRERWTEA EHNRFLEALK LYGRAWQRIE GLIARSMNGE 60 EHKLEKEAIN NGTSPGQAHD IDIPPPRPKR KPNSPYPRKS CLSSETSTRE VQNDKATISN 120 MTNNSTAQMA GDAALEKLTY IQKLQRKEIS EKGSCSEVLN LFREVPSASF SSVNKSSSNH 180 GASRGLEPTK TEVKDVVILE RDSISNGAGK DAKDINDQEM ERLNGIHISS KPDHSHENCL 240 DTSSQQFKPK SNSVETTYVD WSAAKASHYQ MDRNGVTGFQ ATGTEGSHPD QTSDQMGGAS 300 GTMNQCIHPT LPVDPKFDGN AAAQPFPHNY AAFAPMMQCH CNQDAYRSFA NMSSTFSSML 360 VSTLLSNPAI HAAARLAASY WPTVDGNTPD PNQENLSESA QGSHAGSPPN MASIVTATVA 420 AASAWWATQG LLPLFPPPIA FPFVPAPSAP FSTADVQRAQ EKDIDCPMDN AQKELQETRK 480 QDNSEAMKVI VSSETDESGK GEVSLHTELK ISPADKADTK PAAGAETSDV FGNKKKQDRS 540 SCGSNTPSSS DIEADNAPEN QEKANDKAKQ ASCSNSSAGD NNHRRFRSSA STSDSWKEVS 600 EEVVVYQHSA YSHLSYSLNI HHCFSNSHFC CQGRLAFDAL FSRERLPQSF SPPQVEGSKE 660 ISKEEEDEVT TVTVDLNKNA AIIDQELDTA DEPRASFPNE LSNLKLKSRR TGFKPYKRCS 720 VEAKENRVPA SDEVGTKRIR LESEAST |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
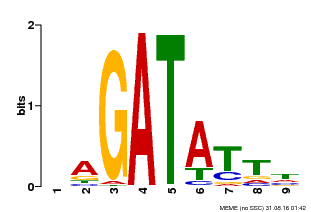
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP004460 | 0.0 | AP004460.2 Oryza sativa Japonica Group genomic DNA, chromosome 8, PAC clone:P0438H08. | |||
| GenBank | AP014964 | 0.0 | AP014964.1 Oryza sativa Japonica Group DNA, chromosome 8, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015649835.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_015649836.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_015649837.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_025875636.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_025875637.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_025875638.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_025875639.1 | 0.0 | protein LHY isoform X1 | ||||
| TrEMBL | A0A0E0QEK1 | 0.0 | A0A0E0QEK1_ORYRU; Uncharacterized protein | ||||
| STRING | ORUFI08G03840.2 | 0.0 | (Oryza rufipogon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 2e-29 | circadian clock associated 1 | ||||



