 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI06G06220.3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 478aa MW: 50713.7 Da PI: 5.0033 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 34.1 | 4.8e-11 | 327 | 372 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
h+++Er RR++i +++ +L+ l+P++ K +Ka++L + ++Y+k Lq
ORUFI06G06220.3 327 HSIAERLRREKISERMKNLQVLVPNS-----NKADKASMLDEIIDYVKFLQ 372
99***********************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 2.36E-16 | 320 | 382 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.65E-12 | 320 | 376 | No hit | No description |
| PROSITE profile | PS50888 | 16.666 | 322 | 371 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 6.7E-16 | 324 | 381 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.4E-8 | 327 | 372 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.1E-13 | 328 | 377 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 478 aa Download sequence Send to blast |
MDYSAGSYMW PGNSGSENYN FVDGSSESYA EEGSLPPSGY FMGAGSDRSL KITENERNPT 60 MLANGCLPYN TQAHPLSGQI LPKGELPNNL LDLQQLQNSS NLRSNSIPPG VLQCNSTSGT 120 FDAKLDTPGL AELPHALSSS IDSNGSDISA FLADVHAVSS APTLCSAFQN VSSFMEPVNL 180 DAFGFQGAQN VAMLNKTSLP NGNPSLFDNA AIASLHDSKE FLNGGSIPSF GTVLQALGAG 240 GLKAAQQEQN IRNIPLPTFT SGSHLAVTDA QGPPLPSKIP PLIHDHNSEY PINHSSDVEP 300 QANSAPGNSA NAKPRTRARR GQATDPHSIA ERLRREKISE RMKNLQVLVP NSNKADKASM 360 LDEIIDYVKF LQLQVKVLSM SRLGAPGAVL PLLRESQTEC HSNPSLSAST ISQGPPDMPD 420 SEDSSAFEQE VVKLMETSIT SAMQYLQNKG LCLMPIALAS AISNQKGMAA AAAIPPEK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
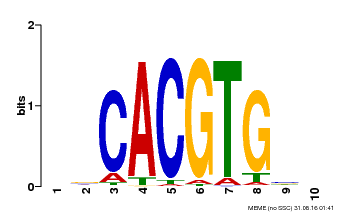
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK065667 | 0.0 | AK065667.1 Oryza sativa Japonica Group cDNA clone:J013042L14, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015644031.1 | 0.0 | transcription factor SPATULA isoform X1 | ||||
| Refseq | XP_025882097.1 | 0.0 | transcription factor SPATULA isoform X1 | ||||
| TrEMBL | A0A0E0PUL1 | 0.0 | A0A0E0PUL1_ORYRU; Uncharacterized protein | ||||
| STRING | ORUFI06G06220.2 | 0.0 | (Oryza rufipogon) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58010.1 | 2e-56 | LJRHL1-like 3 | ||||



