 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI05G01600.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 416aa MW: 47468 Da PI: 8.5554 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 19.5 | 2.6e-06 | 144 | 168 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ C+ s+srk++L+rH+ tH
ORUFI05G01600.1 144 FSCHvdGCPFSYSRKDHLNRHLLTH 168
89*********************99 PP
| |||||||
| 2 | zf-C2H2 | 11.1 | 0.0012 | 173 | 198 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+ Cp C++ F+ k n +rH + H
ORUFI05G01600.1 173 FACPmeGCNRKFTIKGNIQRHVQEmH 198
78*******************98877 PP
| |||||||
| 3 | zf-C2H2 | 20.6 | 1.2e-06 | 210 | 234 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp +Cgk+F+ s L++H +H
ORUFI05G01600.1 210 FICPeeNCGKTFKYASKLQKHEESH 234
78*******************9988 PP
| |||||||
| 4 | zf-C2H2 | 16.8 | 2e-05 | 301 | 325 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
+C+ dC+ sFs ksnL +H + H
ORUFI05G01600.1 301 RCHlkDCKLSFSKKSNLDKHVKAvH 325
7*******************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 15 | 33 | 55 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-9 | 141 | 170 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.74E-6 | 141 | 170 | No hit | No description |
| PROSITE profile | PS50157 | 9.349 | 144 | 173 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0016 | 144 | 168 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 146 | 168 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.15E-5 | 171 | 199 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-6 | 171 | 199 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.866 | 173 | 203 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0019 | 173 | 198 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 175 | 198 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-7 | 206 | 234 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.47E-5 | 207 | 236 | No hit | No description |
| PROSITE profile | PS50157 | 11.24 | 210 | 234 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0018 | 210 | 234 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 212 | 234 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 7.8 | 242 | 267 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 244 | 267 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 13 | 270 | 291 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 2.48E-7 | 281 | 342 | No hit | No description |
| PROSITE profile | PS50157 | 11.967 | 300 | 330 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.031 | 300 | 325 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.4E-5 | 301 | 326 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 302 | 325 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.6E-6 | 327 | 354 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 4.7 | 331 | 357 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.974 | 331 | 362 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 333 | 357 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 416 aa Download sequence Send to blast |
MGSVELGAEE REVAGGEGGS KGAAPPARDI RRYKCDFCSV VRSKKGLIRA HVLEHHKDEV 60 DDLDDYLGRG GGETCKEMDH DCKHWIPENS KMREGLEWTS TRMGRPFWEA PSRMPLSRSS 120 SGWGDSPEKL HQECLVHTLF QRPFSCHVDG CPFSYSRKDH LNRHLLTHQG KLFACPMEGC 180 NRKFTIKGNI QRHVQEMHKD GSPCESKKEF ICPEENCGKT FKYASKLQKH EESHVKLDYS 240 EVICCEPGCM KAFTNLECLK AHNKSCHRHV VCDVCGTKQL KKNFKRHQRM HEGSCVTERV 300 RCHLKDCKLS FSKKSNLDKH VKAVHEQKRP FVCGFSGCGK SFSYKHVRDN HEKSRAHVYV 360 QANFEEIDGE RPRQAGGRKR KAIPVESLMR KRVAAPDDDA PACDDGTEYL RWLLSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 2e-13 | 141 | 264 | 40 | 157 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 2e-13 | 141 | 264 | 40 | 157 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
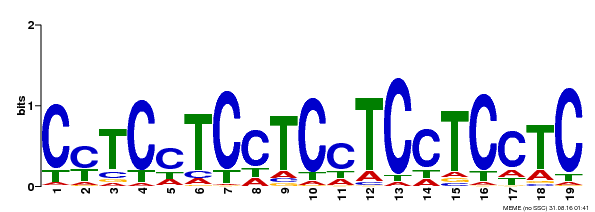
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK101631 | 0.0 | AK101631.1 Oryza sativa Japonica Group cDNA clone:J033054O17, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015637660.1 | 0.0 | transcription factor IIIA isoform X1 | ||||
| TrEMBL | A0A0E0PGU4 | 0.0 | A0A0E0PGU4_ORYRU; Uncharacterized protein | ||||
| STRING | ORUFI05G01600.1 | 0.0 | (Oryza rufipogon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 7e-94 | transcription factor IIIA | ||||



