 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI02G29230.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 224aa MW: 23916.5 Da PI: 4.9599 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 57.2 | 4.2e-18 | 53 | 103 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ +kGVr++ grWv+e+r+p g ++r++lg+f tae+Aa+a++ a+++l+g
ORUFI02G29230.1 53 PVFKGVRRRN-PGRWVCEVREP--HG-KQRIWLGTFETAEMAARAHDVAALALRG 103
579****887.9******9996..33.5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.9E-13 | 53 | 103 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 6.54E-21 | 54 | 112 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 4.5E-28 | 54 | 117 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 21.667 | 54 | 111 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.9E-30 | 54 | 113 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 8.6E-8 | 55 | 66 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.05E-20 | 55 | 113 | No hit | No description |
| PRINTS | PR00367 | 8.6E-8 | 77 | 93 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 224 aa Download sequence Send to blast |
MDVSAALSSD YSSGTPSPVA ADADDGSSAY MTVSSAPPKR RAGRTKFKET RHPVFKGVRR 60 RNPGRWVCEV REPHGKQRIW LGTFETAEMA ARAHDVAALA LRGRAACLNF ADSPRRLRVP 120 PIGASHDDIR RAAAEAAEAF RPPPDESNAA TEVAAAASGA TNSNAEQFAS HPYYEVMDDG 180 LDLGMQGYLD MAQGMLIDPP PMAGDPAVGG GEDDNDGEVQ LWSY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 6e-14 | 52 | 110 | 12 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00644 | PBM | Transfer from LOC_Os02g45450 | Download |
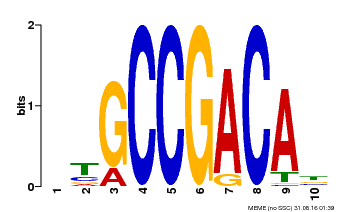
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK060550 | 0.0 | AK060550.1 Oryza sativa Japonica Group cDNA clone:001-021-H10, full insert sequence. | |||
| GenBank | AK106041 | 0.0 | AK106041.1 Oryza sativa Japonica Group cDNA clone:001-206-E01, full insert sequence. | |||
| GenBank | AP005775 | 0.0 | AP005775.3 Oryza sativa Japonica Group genomic DNA, chromosome 2, BAC clone:OSJNBb0005A04. | |||
| GenBank | AP006060 | 0.0 | AP006060.3 Oryza sativa Japonica Group genomic DNA, chromosome 2, PAC clone:P0135D07. | |||
| GenBank | AP014958 | 0.0 | AP014958.1 Oryza sativa Japonica Group DNA, chromosome 2, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015624757.1 | 1e-162 | dehydration-responsive element-binding protein 1G | ||||
| Swissprot | Q6EP77 | 1e-163 | DRE1G_ORYSJ; Dehydration-responsive element-binding protein 1G | ||||
| TrEMBL | A0A0D9YWE3 | 1e-162 | A0A0D9YWE3_9ORYZ; Uncharacterized protein | ||||
| TrEMBL | A0A0E0NJ65 | 1e-162 | A0A0E0NJ65_ORYRU; Uncharacterized protein | ||||
| STRING | OGLUM02G28340.1 | 1e-163 | (Oryza glumipatula) | ||||
| STRING | ORUFI02G29230.1 | 1e-163 | (Oryza rufipogon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP196 | 36 | 261 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G25470.1 | 6e-35 | C-repeat/DRE binding factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



