 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORUFI01G09400.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 786aa MW: 84970.6 Da PI: 9.7203 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 38.9 | 1.6e-12 | 632 | 678 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
ORUFI01G09400.1 632 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAITYITDLQ 678
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 8.6E-50 | 229 | 414 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.831 | 628 | 677 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.33E-15 | 631 | 682 | No hit | No description |
| SuperFamily | SSF47459 | 1.57E-18 | 631 | 694 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 5.1E-10 | 632 | 678 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 8.9E-18 | 632 | 691 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.9E-16 | 634 | 683 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 786 aa Download sequence Send to blast |
MNKPPARKAE KSPWVGDRAG SPAPNEPRTG SARLGSGSNP TGVEKAKAPS FSFHTSPPLF 60 FSLSTSQVAS LLRPRNPLAN QRNLRHHHHH HRLRLRLRLL RARRIRGTLA LAWGIRASPS 120 PGGGGCSDSA SGGWGGPPLS SWGGAEETPP VLDSVAAAPI RAREPQHGIM VMKMEADEDG 180 ANGGTGGTWT DEDRALTASV LGTDAFAYLT KGGGAISEGL VAASLPVDLQ NRLQELVESD 240 RPGAGWNYAI FWQLSRTKSG DLVLGWGDGS CREPHDGEMG PAASAGSDEA KQRMRKRVLQ 300 RLHSAFGGVD EEDYAPGIDQ VTDTEMFFLA SMYFAFPRRA GGPGQVFAAG VPLWIPNTER 360 NVFPANYCYR GYLANAAGFR TIVLVPFETG VLELGSMQQV AESSDTLQTI RSVFAGAIGN 420 KAGVQRHEGS GPTDKSPGLA KIFGKDLNLG RPSAGPGTGV SEADERSWEQ RTGGGSSLLP 480 NVQRGLQNFT WSQARGLNSH QQKFGNGILI VSNEATPRNN GVVDSSTATQ FQLQKAPPLQ 540 KLPQLQKSHQ LVKPQQLVSQ QQLQPQAPRQ IDFSAGTSSK PGVLTKKPAG IDGDGAEVDG 600 LCKDEGPPPA LEDRRPRKRG RKPANGREEP LNHVEAERQR REKLNQRFYA LRAVVPNISK 660 MDKASLLGDA ITYITDLQKK LKEMEVERER LIESGMIDPR DRTPRPEVDI QVVQDEVLVR 720 VMSPMESHPV RAIFQAFEEA EVHAGESKIT SNNGTAVHSF IIKCPGAEQQ TREKVIAAMS 780 RVMNSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 2e-26 | 626 | 688 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 613 | 621 | RRPRKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
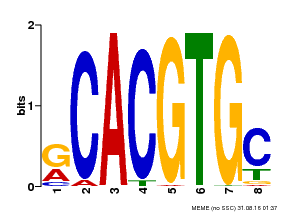
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CT833046 | 0.0 | CT833046.1 Oryza sativa (indica cultivar-group) cDNA clone:OSIGCRA221E23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015621308.1 | 0.0 | transcription factor bHLH13 | ||||
| TrEMBL | A0A0E0MTM9 | 0.0 | A0A0E0MTM9_ORYRU; Uncharacterized protein | ||||
| STRING | ORUFI01G09400.1 | 0.0 | (Oryza rufipogon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 4e-70 | bHLH family protein | ||||



