 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC12G10470.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 271aa MW: 29040.6 Da PI: 9.6516 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 144.1 | 7.7e-45 | 17 | 151 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdL.pkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevl 93
lppGfrF+Ptdeelvv+yL++++ +++l+ + i++v+++ ++P+dL p +e+e yfF++++ ky +g+r+nrat +gyWkatgk+k+v
OPUNC12G10470.1 17 LPPGFRFRPTDEELVVHYLRRRALDSPLPPAVDIPDVRLLAHDPSDLlP--PGWSEQERYFFTCKEAKYVKGRRANRATGAGYWKATGKEKPVA 108
79*************************988556**************43..3447899************************************ PP
NAM 94 sk........kgelvglkktLvfykgrapkgektdWvmheyrl 128
++ +vg+k++Lvfy+g+ p+g+ktdWvmheyrl
OPUNC12G10470.1 109 VSvpaaprsqAAVVVGMKRSLVFYRGKPPTGKKTDWVMHEYRL 151
99888886555555***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 6.28E-50 | 7 | 181 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 48.998 | 17 | 181 | IPR003441 | NAC domain |
| Pfam | PF02365 | 9.2E-27 | 18 | 151 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 271 aa Download sequence Send to blast |
METTAKECAA RRAAAALPPG FRFRPTDEEL VVHYLRRRAL DSPLPPAVDI PDVRLLAHDP 60 SDLLPPGWSE QERYFFTCKE AKYVKGRRAN RATGAGYWKA TGKEKPVAVS VPAAPRSQAA 120 VVVGMKRSLV FYRGKPPTGK KTDWVMHEYR LAGAGLAPCR RAADHPARPA EGWVLCRVFR 180 KKGSSAASTA SPTADADEDD ATTVRADDGG DGAVTAAGVR FIDFFARADA RRRRAASPVS 240 SSCVTDASAE HCREQETTSR GGGAGAGDAS D |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 8e-38 | 13 | 184 | 11 | 171 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor that negatively regulates the expression of genes involved in xylem vessel formation. Represses the transcriptional activation activity of NAC030/VND7, which regulates protoxylem vessel differentiation by promoting immature xylem vessel-specific genes expression (PubMed:20388856). Transcriptional activator that regulates the COLD-REGULATED (COR15A and COR15B) and RESPONSIVE TO DEHYDRATION (LTI78/RD29A and LTI65/RD29B) genes by binding directly to their promoters. Mediates signaling crosstalk between salt stress response and leaf aging process (PubMed:21673078). May play a role in DNA replication of mungbean yellow mosaic virus (PubMed:24442717). {ECO:0000269|PubMed:20388856, ECO:0000269|PubMed:21673078, ECO:0000269|PubMed:24442717}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00507 | DAP | Transfer from AT5G13180 | Download |
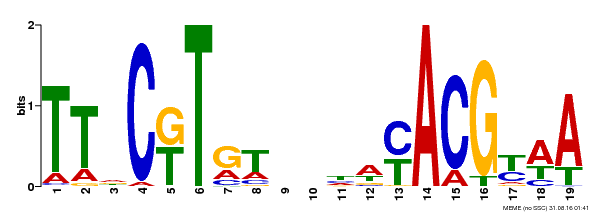
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA) and salt stress. {ECO:0000269|PubMed:21673078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK243244 | 0.0 | AK243244.1 Oryza sativa Japonica Group cDNA, clone: J100046N20, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015618303.1 | 1e-135 | NAC domain-containing protein 83 | ||||
| Swissprot | Q9FY93 | 1e-52 | NAC83_ARATH; NAC domain-containing protein 83 | ||||
| TrEMBL | A0A0E0MMB1 | 0.0 | A0A0E0MMB1_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC12G10470.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6317 | 32 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13180.1 | 1e-53 | NAC domain containing protein 83 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



