 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC11G01450.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 631aa MW: 69208.3 Da PI: 4.5653 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.8 | 1.8e-16 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++++G+g+W++++ g+ R+ k+c++rw +yl
OPUNC11G01450.1 14 KGPWTPEEDIILVSYIQEHGPGNWRSVPINTGLMRCSKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 45.2 | 2.2e-14 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T+ E+ ++v++ +lG++ W++Ia++++ Rt++++k++w+++l
OPUNC11G01450.1 67 RGNFTAHEEGIIVHLQSLLGNR-WAAIASYLP-QRTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-23 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 25.517 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.73E-29 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.4E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.0E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.12E-11 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-23 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.1E-13 | 66 | 114 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 18.443 | 66 | 116 | IPR017930 | Myb domain |
| Pfam | PF00249 | 4.2E-12 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.01E-9 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 631 aa Download sequence Send to blast |
MGRPPCCDKE GIKKGPWTPE EDIILVSYIQ EHGPGNWRSV PINTGLMRCS KSCRLRWTNY 60 LRPGIKRGNF TAHEEGIIVH LQSLLGNRWA AIASYLPQRT DNDIKNYWNT HLKKKLAANS 120 NSSSNRHPIF ADATFPSAAT GSHTASSNSD VNQMAAIARR SPFADCPSSS YASSMDNISK 180 LLDGFMKTNS PSPPPPPLHH YDGGCYDDVK PTVDVGNPLL SFDCMSGADL DCFDVHQQQQ 240 QPASFMEYGG YGGGYGDESK QQLMNQATPP LSSIEKWLFD EAAAEQVADL MDLSDGRQNP 300 FDSSNGSDEI IELPHVPDSA PSCLLPRINL FDLAVNQNMA HDNALQETWT PLSYFSARRH 360 RNYGNLYVRH STILHHNSFK LEKDEISEND AHNSQSDGDA KQEGNTSKLF GSLEAHIGEE 420 IKILGAAISD VGVLEVNSGK DSGNQNADFS YDISSSPIQK SRQSTFEAKD AVHAGIEQLT 480 LCSPYKVNNF EAHIVEADSI YEFNSLFKCR MEEVLVQSIS ESSISQPLTV KLEDESSEPL 540 SSDSGTGTHF IDGSSVEDSD PQFAQLNDEA LVCATSNATC RNESIEEKSS EALLVGNKDY 600 SELPNELLKS GDPQTADISE IQVQVIEATG H |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 7e-26 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 7e-26 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00132 | DAP | Transfer from AT1G08810 | Download |
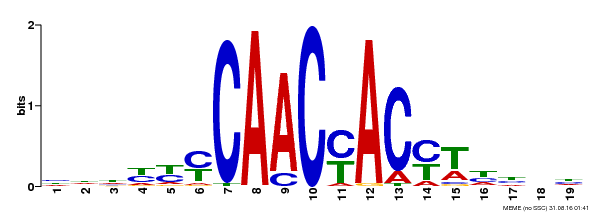
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK242802 | 0.0 | AK242802.1 Oryza sativa Japonica Group cDNA, clone: J090059L09, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015618674.1 | 0.0 | uncharacterized protein LOC4351393 | ||||
| Refseq | XP_025877882.1 | 0.0 | uncharacterized protein LOC4351393 | ||||
| Refseq | XP_025877883.1 | 0.0 | uncharacterized protein LOC4351393 | ||||
| TrEMBL | A0A0E0MBY8 | 0.0 | A0A0E0MBY8_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC11G01450.1 | 0.0 | (Oryza punctata) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08810.1 | 1e-83 | myb domain protein 60 | ||||



