 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC06G21190.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 424aa MW: 44654 Da PI: 9.1311 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.1 | 3.9e-41 | 176 | 252 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq+egC+adls ak+yhrrhkvCe h+ka+vv++sg++qrfCqqCsrfh l+efDe+krsCr+rLa+hn+rrrk++
OPUNC06G21190.1 176 RCQAEGCKADLSGAKHYHRRHKVCEYHAKASVVAASGKQQRFCQQCSRFHVLTEFDEAKRSCRKRLAEHNRRRRKPA 252
6**************************************************************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.6E-35 | 169 | 238 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.547 | 174 | 251 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 9.16E-39 | 175 | 254 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.2E-31 | 177 | 250 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 424 aa Download sequence Send to blast |
MMSGRMNAAG DESSFPFGAM QAPGGTTYVG FDHGAAAVAA AAAAAQRAGM QQQHHHHHHH 60 MYDGLDFAAA MQFGGQEAPQ LLALPPSMAP MAAPPPMPLP LQMPVPMPGD VYPALGIVKR 120 EGGGGQDASG RIGLNLGRRT YFSPGDMLAV DRLLMRSRLG GVFGLGFGGA HHQPPRCQAE 180 GCKADLSGAK HYHRRHKVCE YHAKASVVAA SGKQQRFCQQ CSRFHVLTEF DEAKRSCRKR 240 LAEHNRRRRK PATTTTTAAA AKDAAAAVAA GKKPSGATTS YTGDSKNVVV SMSAAKSPIS 300 SNTSVISCLA EQGKQAAARP TALTLGGAPP PHESSAQLGA VLHAHHHHHH HHQQDHMQVS 360 SLVHINGGGG SNNILSCSSV CSSALPSTAT NGEVSDQNND NSHNNGSNNN NNNNNMHLFE 420 VDFM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-30 | 177 | 250 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 233 | 250 | KRSCRKRLAEHNRRRRKP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
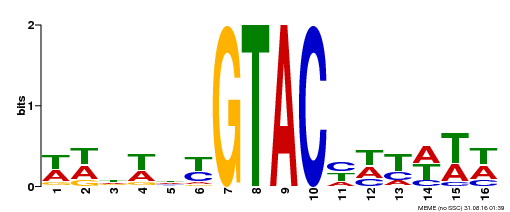
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012614 | 0.0 | CP012614.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 6 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015642406.1 | 1e-166 | squamosa promoter-binding-like protein 10 | ||||
| Swissprot | A2YFT9 | 1e-167 | SPL10_ORYSI; Squamosa promoter-binding-like protein 10 | ||||
| Swissprot | Q0DAE8 | 1e-167 | SPL10_ORYSJ; Squamosa promoter-binding-like protein 10 | ||||
| TrEMBL | A0A0E0LE87 | 0.0 | A0A0E0LE87_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC06G21190.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2375 | 35 | 93 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 2e-49 | squamosa promoter binding protein-like 8 | ||||



