 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC05G11000.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 260aa MW: 27001.1 Da PI: 9.2888 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.9 | 6.1e-19 | 45 | 94 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+WvAeIr+p++ r r +lg+f tae+Aa+a+++a++ l+g
OPUNC05G11000.1 45 RYRGVRQRS-WGKWVAEIREPRK---RSRKWLGTFATAEDAARAYDRAALLLYG 94
59****999.**********954...5**********************98877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 1.2E-12 | 45 | 93 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 6.54E-21 | 45 | 102 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 2.1E-36 | 45 | 108 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.379 | 45 | 102 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.27E-17 | 45 | 100 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.1E-29 | 45 | 101 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.3E-10 | 46 | 57 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.3E-10 | 68 | 84 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006970 | Biological Process | response to osmotic stress | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0009749 | Biological Process | response to glucose | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010353 | Biological Process | response to trehalose | ||||
| GO:0010449 | Biological Process | root meristem growth | ||||
| GO:0010896 | Biological Process | regulation of triglyceride catabolic process | ||||
| GO:0031930 | Biological Process | mitochondria-nucleus signaling pathway | ||||
| GO:0032880 | Biological Process | regulation of protein localization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 260 aa Download sequence Send to blast |
MQPSVDAATA AVAAPAETAS SSGGGGRSTA KKAAGKGGPE NGKFRYRGVR QRSWGKWVAE 60 IREPRKRSRK WLGTFATAED AARAYDRAAL LLYGPRAHLN LTAPPPLPPA PSSDSAAAAA 120 SSSSAASSTS AAPPLRPLLP RPPHLHPAFH HQQFHHHLLQ QQPPPPPPLY YATTASTSTV 180 TTTTTTTTAP PPQLTAAAPA VLVAAAVSST AETQAVAAPE DAAAAAAAEV AWGYHGGDEE 240 DYAAALLWSE PDPWFDLFLK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-16 | 46 | 101 | 6 | 62 | ATERF1 |
| 3gcc_A | 1e-16 | 46 | 101 | 6 | 62 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription regulator that binds to cis-responsive elements of genes involved in stress response. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00674 | SELEX | Transfer from GRMZM2G093595 | Download |
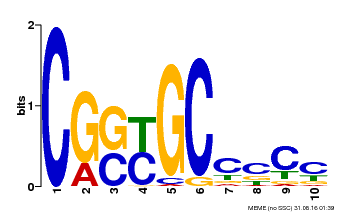
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC136221 | 0.0 | AC136221.2 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone OSJNBa0077J17, complete sequence. | |||
| GenBank | AP014961 | 0.0 | AP014961.1 Oryza sativa Japonica Group DNA, chromosome 5, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003568646.1 | 1e-63 | ethylene-responsive transcription factor ABI4 | ||||
| Swissprot | C7J2Z1 | 2e-53 | ABI4_ORYSJ; Ethylene-responsive transcription factor ABI4 | ||||
| TrEMBL | A0A0E0L1C0 | 1e-180 | A0A0E0L1C0_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC05G11000.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP13303 | 26 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40220.1 | 2e-18 | ERF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



