 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI12G12160.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | B3 | ||||||||
| Protein Properties | Length: 1674aa MW: 190554 Da PI: 9.2696 | ||||||||
| Description | B3 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 45.1 | 1.8e-14 | 59 | 133 | 18 | 98 |
--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE- CS
B3 18 lpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfr 98
+p kfa+ + g+ s ++le ++g++++v++ k +++ vl++GW F+++ +Lk+ D++vF+++g+s+f +v++f+
OMERI12G12160.1 59 VPTKFANIIRGQ--ISEVVKLEVPNGKTYDVQV--AKEQNELVLRSGWGAFARDYELKQCDILVFTYSGSSRF--KVRIFN 133
799999666766..6779***************..*********************************99999..999987 PP
| |||||||
| 2 | B3 | 49.5 | 7.5e-16 | 183 | 263 | 12 | 98 |
HTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE- CS
B3 12 ksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfr 98
+ +++pk fa++ +g+ s +le ++gr+++v++ k ++ v+++GW+ F++a +L++gD++vF ++g+s f +v++f+
OMERI12G12160.1 183 LDKPVTIPKMFAKNVQGQ--ISGVAKLEVPDGRTYDVEI--AKEHNELVFRSGWEVFASAYELEQGDILVFAYSGNSHF--KVQIFN 263
555689******888888..6669***************..**********************************9999..999987 PP
| |||||||
| 3 | B3 | 54.8 | 1.7e-17 | 370 | 461 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
ffkv+++ + k l +pkkf+++ gk + +++le ++g+ ++v++ + +++ vl++GW +F++ +Lk gD +vF ++g+s f +v
OMERI12G12160.1 370 FFKVMMSVSDIKDE-LAIPKKFTANVRGK--IPEQVRLEVSDGKMYNVQV--TEEQDELVLRSGWANFASTYQLKHGDLLVFIHSGHSHF--KV 456
99***999999987.********888666..3446***************..********************************999999..99 PP
EEE-S CS
B3 95 kvfrk 99
+f+
OMERI12G12160.1 457 LIFDP 461
99885 PP
| |||||||
| 4 | B3 | 35.6 | 1.6e-11 | 587 | 666 | 12 | 92 |
HTT-EE--HHH.HTT---..--SEEEEEETTS.-EEEEEE..EEETTE.EEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE.. CS
B3 12 ksgrlvlpkkfaeehggkkeesktltledesg.rsWevkliyrkksgr.yvltkGWkeFvkangLkegDfvvFkldgr.sefel 92
+sg lv++k++a ++ ++ +++t++l ++ g ++W++ l + + y l++GW +F + n L+egD++vF+ +++ ++ l
OMERI12G12160.1 587 ASGSLVFSKDYAVRYLLD--QNRTIKLCQSGGsKTWDISL--DMDTDDlYALSTGWLDFFRGNLLQEGDICVFEASKSkRGVAL 666
45679**********888..4558888887777*******..8888888*************************8876444444 PP
| |||||||
| 5 | B3 | 42.1 | 1.5e-13 | 728 | 826 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefel 92
f++v+ +s+ ++++ l+++ k+a+++ e++ ++l+ e+ + + l+++ + + + l kGW +Fv++n+L++ D+++F+l+++ ++ ++
OMERI12G12160.1 728 FLSVMRSSNCTRQSSLCFSVKYASKYLPH--EDQNMRLRLpETKYKCKAALRVDTSTNLHKLLKGWGKFVNDNKLEVHDICLFQLMKNkKKLTM 819
7889999999999*************533..3446777773455788888877999999***************************999***** PP
EEEEE-S CS
B3 93 vvkvfrk 99
+v+++rk
OMERI12G12160.1 820 TVHIIRK 826
****997 PP
| |||||||
| 6 | B3 | 57.2 | 3e-18 | 866 | 953 | 1 | 97 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
f+kv+ +k+g +++p kfa+++gg+ s t++le+++g+++ev++ k + vl++GW++F++a +L+ gD++vF +g+s f +v
OMERI12G12160.1 866 FIKVM--ITDFKNG-VTIPAKFARNFGGQM--SGTVKLETRNGKTYEVQV--AKELNNLVLRSGWERFASAYELEKGDILVFIQSGNSHF--KV 950
67777..6678887.********9999994..559***************..*******************************8888888..66 PP
EEE CS
B3 95 kvf 97
++
OMERI12G12160.1 951 WIY 953
665 PP
| |||||||
| 7 | B3 | 40.2 | 5.9e-13 | 1082 | 1163 | 8 | 93 |
-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..E CS
B3 8 sdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelv 93
+ + + g+l ++k++a ++ ++k+e t++l + ++ +W++ l ++ +++y l++GW +F + n+L+egD++vF+ ++++++ v
OMERI12G12160.1 1082 KASATDGLLAISKDYAVSYLLDKNE--TIKLCHsGRSLTWDIGLDIDT-DDQYALSTGWLDFIRNNHLQEGDICVFEASKNKRG--V 1163
55677899***********888767..56665537889******4333.346*************************8887666..3 PP
| |||||||
| 8 | B3 | 37.5 | 4.1e-12 | 1240 | 1320 | 13 | 94 |
TT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE..EE CS
B3 13 sgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefelvv 94
s+ ++++ k+a+++ + e++ +++ + +s++ k ++ + + + l++GW +Fv++n+Lk gD+++F+l+++ ++ +++v
OMERI12G12160.1 1240 SRYMCFSAKYAAKYL-PRENN-KIMRLRLPKKSYKYKAVFKINNKVHKLGGGWGKFVDDNKLKLGDICLFQLMKDkKKLTMMV 1320
556788889999993.33334.455555566777777777999999***************************9867776666 PP
| |||||||
| 9 | B3 | 42.8 | 9.5e-14 | 1358 | 1434 | 14 | 94 |
T-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 14 grlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
+ +++p k ae ++g+ ++ l+++sg++W+v + +k ++ +l +GW++F+ka++L+e+D + F+ +g+ ++++ +
OMERI12G12160.1 1358 HGISIPEKVAEIFSGQITK--GFNLKSPSGETWRVGI--EKVADELILMSGWEDFAKAHELQENDLLFFTCNGHGNGSCSF 1434
459*******777777555..59999***********..*******************************77665554443 PP
| |||||||
| 10 | B3 | 44.4 | 3e-14 | 1567 | 1663 | 1 | 97 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE...EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliy.rkksgryvltkGWkeFvkangLkegDfvvFkldgr.se 89
f+ vl +++v ++++l++p +fa++h + +s ++ l + ++W+vk+ y +++ +++ W +F ++n+L+eg +++F+l++ ++
OMERI12G12160.1 1567 FVAVLQTAHVRRRNILIVPTRFAADHLER--KSHDILLIRpNRKQKWSVKYYYlSNTTRGFNCH-RWIKFIRENRLREGNVCIFELMKGaRR 1655
7889999*******************544..45567776636779******8865555555544.8*******************9976688 PP
E..EEEEE CS
B3 90 felvvkvf 97
+++v+v+
OMERI12G12160.1 1656 PTMTVHVI 1663
88888775 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 2.4E-18 | 23 | 137 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 2.8E-6 | 55 | 135 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 2.55E-18 | 55 | 137 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 11.293 | 59 | 135 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 9.3E-12 | 59 | 133 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 7.69E-17 | 59 | 133 | No hit | No description |
| PROSITE profile | PS50863 | 12.224 | 172 | 265 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 8.3E-9 | 173 | 265 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 9.1E-19 | 175 | 267 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.39E-18 | 176 | 267 | IPR015300 | DNA-binding pseudobarrel domain |
| Pfam | PF02362 | 1.0E-13 | 182 | 263 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 8.87E-17 | 188 | 263 | No hit | No description |
| SuperFamily | SSF101936 | 1.55E-20 | 363 | 462 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 7.4E-21 | 365 | 462 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.92E-17 | 368 | 460 | No hit | No description |
| PROSITE profile | PS50863 | 12.76 | 369 | 462 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 6.0E-15 | 370 | 460 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.2E-10 | 370 | 462 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 1.7E-13 | 570 | 673 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.31E-14 | 571 | 674 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 11.124 | 576 | 674 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.7E-4 | 578 | 674 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 9.1E-9 | 587 | 666 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 1.05E-9 | 588 | 672 | No hit | No description |
| Gene3D | G3DSA:2.40.330.10 | 7.0E-17 | 720 | 827 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 5.49E-19 | 721 | 829 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.52E-15 | 726 | 825 | No hit | No description |
| Pfam | PF02362 | 2.9E-12 | 728 | 826 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 7.5E-8 | 728 | 827 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 9.601 | 728 | 827 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 1.65E-21 | 860 | 958 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.6E-21 | 862 | 958 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 13.084 | 863 | 956 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 1.91E-19 | 864 | 954 | No hit | No description |
| Pfam | PF02362 | 3.5E-16 | 866 | 952 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.4E-15 | 866 | 956 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 2.3E-15 | 1069 | 1167 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.51E-16 | 1069 | 1168 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.111 | 1075 | 1170 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.4E-5 | 1075 | 1173 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.0E-9 | 1083 | 1162 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 4.25E-10 | 1090 | 1155 | No hit | No description |
| Gene3D | G3DSA:2.40.330.10 | 9.7E-16 | 1219 | 1325 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.0E-17 | 1220 | 1326 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.04E-15 | 1225 | 1320 | No hit | No description |
| SMART | SM01019 | 0.0015 | 1227 | 1324 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 10.165 | 1228 | 1324 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.1E-10 | 1239 | 1320 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 5.3E-17 | 1341 | 1444 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.167 | 1345 | 1442 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 2.90E-13 | 1347 | 1440 | No hit | No description |
| Gene3D | G3DSA:2.40.330.10 | 3.3E-16 | 1348 | 1444 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 3.3E-7 | 1348 | 1442 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 5.4E-11 | 1357 | 1432 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 2.0E-17 | 1560 | 1665 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 9.42E-20 | 1561 | 1661 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.34E-17 | 1565 | 1661 | No hit | No description |
| Pfam | PF02362 | 3.9E-12 | 1567 | 1663 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 6.0E-10 | 1567 | 1667 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.155 | 1567 | 1666 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005773 | Cellular Component | vacuole | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1674 aa Download sequence Send to blast |
MRKSFTTCKE CIFYHYWNHM GDQKRSFINV MIGDFVPNAL HEVHSKLDVG LIWLFLQAVP 60 TKFANIIRGQ ISEVVKLEVP NGKTYDVQVA KEQNELVLRS GWGAFARDYE LKQCDILVFT 120 YSGSSRFKVR IFNPSGCEKE LSCVMMNSTP CGHEGSMPYH DNHVQSPSER SSQHATVLPP 180 VLLDKPVTIP KMFAKNVQGQ ISGVAKLEVP DGRTYDVEIA KEHNELVFRS GWEVFASAYE 240 LEQGDILVFA YSGNSHFKVQ IFNPSNCEKE LSCVMMNRSI SDDNHRQSPS WESLVPASRR 300 RRSPPACRLT APLPASQHAA YLQAAPPPAT AVRINVALKF IIVMNKSGTT CMDCITNHYW 360 LHMDDRERYF FKVMMSVSDI KDELAIPKKF TANVRGKIPE QVRLEVSDGK MYNVQVTEEQ 420 DELVLRSGWA NFASTYQLKH GDLLVFIHSG HSHFKVLIFD PSYTEKEFSC VVTDSTSHVH 480 ERSISHDNHL QSPRSVILGK NYSLCSSRKR SRMNPADYPS QRPDVPSSED IKDPMSSGGL 540 QKSKKSCYVL PMLYSMTSAQ EAEVLALEKK IQPQIPLYIT AMDISVASGS LVFSKDYAVR 600 YLLDQNRTIK LCQSGGSKTW DISLDMDTDD LYALSTGWLD FFRGNLLQEG DICVFEASKS 660 KRGVALTFHP FKESHCPKSS EYTLSTKSPT RRVPKRDYFA TNLTNLTDQQ ERKVKNKIKS 720 IQSDIPIFLS VMRSSNCTRQ SSLCFSVKYA SKYLPHEDQN MRLRLPETKY KCKAALRVDT 780 STNLHKLLKG WGKFVNDNKL EVHDICLFQL MKNKKKLTMT VHIIRKGECS GSLCCRCCGM 840 EKSHRVCKNC VANHYWLHMD NHGKSFIKVM ITDFKNGVTI PAKFARNFGG QMSGTVKLET 900 RNGKTYEVQV AKELNNLVLR SGWERFASAY ELEKGDILVF IQSGNSHFKV WIYDPSACEK 960 ELPCIITEQL PRVQQRSISH DNHTRLKRNA KSAKLYVDSS GHTKETSEIN PASSPSWKPT 1020 ERVPFSEELD EPVDLANVQK ATKSFYSLPR MCNLTSAQKA EVDALEKRIK PQIPFYITVI 1080 DKASATDGLL AISKDYAVSY LLDKNETIKL CHSGRSLTWD IGLDIDTDDQ YALSTGWLDF 1140 IRNNHLQEGD ICVFEASKNK RGVALIFHPL KQSHHPKPPG CVPSTKFPRH GVSKPNYIVS 1200 RFTTLSGQLK IKVEAKVQAI QSEVPIFVAV MRESFIRGRS RYMCFSAKYA AKYLPRENNK 1260 IMRLRLPKKS YKYKAVFKIN NKVHKLGGGW GKFVDDNKLK LGDICLFQLM KDKKKLTMMV 1320 MGEKGCESCR KWQEHYYKEH MDVSRIRFFR LMTGDFAHGI SIPEKVAEIF SGQITKGFNL 1380 KSPSGETWRV GIEKVADELI LMSGWEDFAK AHELQENDLL FFTCNGHGNG SCSFDVLIFD 1440 ASGCEKVSCF FTGKKNSYMC KNFNSIGGQI AGKYLSSDSE DTSTPSVLIG SPHKASTSKK 1500 LSGKTKTNPR KEPEDPNCSH WHVIEEKNTD DDEHADYHYT RFANYLTGEE RDEIFSLVSL 1560 QPGNPVFVAV LQTAHVRRRN ILIVPTRFAA DHLERKSHDI LLIRPNRKQK WSVKYYYLSN 1620 TTRGFNCHRW IKFIRENRLR EGNVCIFELM KGARRPTMTV HVICKADNRF VLLS |
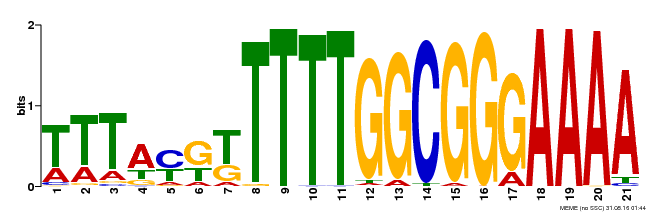
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00468 | DAP | Transfer from AT4G33280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK241993 | 0.0 | AK241993.1 Oryza sativa Japonica Group cDNA, clone: J075099J10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025878051.1 | 0.0 | B3 domain-containing protein LOC_Os12g40080 | ||||
| Swissprot | Q2QMT6 | 0.0 | Y1208_ORYSJ; B3 domain-containing protein LOC_Os12g40080 | ||||
| TrEMBL | A0A0E0FDM3 | 0.0 | A0A0E0FDM3_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI12G12100.1 | 0.0 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP562 | 34 | 179 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G33280.1 | 9e-27 | B3 family protein | ||||



