 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI08G03470.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 746aa MW: 81393 Da PI: 6.5659 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47 | 6e-15 | 51 | 95 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E+ ++++a k++G W +I +++g ++t+ q++s+ qk+
OMERI08G03470.2 51 RERWTEAEHNRFLEALKLYGRA-WQRIEEHVG-TKTAVQIRSHAQKF 95
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.99E-16 | 45 | 101 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.593 | 46 | 100 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.6E-16 | 49 | 98 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.6E-12 | 50 | 98 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.8E-12 | 51 | 94 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.1E-8 | 51 | 91 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.33E-8 | 53 | 96 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 746 aa Download sequence Send to blast |
MQLLSRRRRG RATARLGIRS VVVAVVLCSI NSSGEEAVVK VRKPYTITKQ RERWTEAEHN 60 RFLEALKLYG RAWQRIEEHV GTKTAVQIRS HAQKFFTKLE KEAINNGTSP GQAHDIDIPP 120 PRPKRKPNSP YPRKSCLSSE TSTREVQNDK ATISNMTNNS TAQMAGDAAL EKLQRKEISE 180 KGSCSEVLNL FREVPSASFS SVNKSSSNHG ASRGLEPTKT EVKDVVILER DSISNGAGKD 240 AKDINDQEME RLNGIHISSK PDHSHENCLD TSSQQFKPKS NSVETTYVDW SAAKASHYQM 300 DRNGVTGFQA TGTEGSHPDQ TSDQMGGASG TMNQCIHPTL PVDPKFDGNA AAQPFPHNYA 360 AFSPMMQCHC NQDAYRSFAN MSSTFSSMLV STLLSNPAIH AAARLAASYW PTVDGNTPDP 420 NQENLSESAQ GSHAGSPPNM ASIVAATVAA ASAWWATQGL LPLFPPPIAF PFVPAPSAPF 480 STADVQRAQE KDIDCPMDNA QKELQETRKQ DNSEAMKVIV SSETDESGKG EVSLHTELKI 540 SPADKADTKP AAGAETSDVF GNKKKQDRSS CGSNTPSSSD IEADNAPENQ EKANDKAKQA 600 SCSNSSAGDN NHRRFRSSAS TSDSWKEVSE EGRLAFDALF SRERLPQSFS PPQVEGSKEI 660 SKEEEDEVTT VTVDLNKNAA IIDQELDTAD EPRASFPNEL SNLKLKSRRT GFKPYKRCSV 720 EAKENRVPAS DEVGTKRIRL ESEAST |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the circadian clock. Binds to the promoter region of APRR1/TOC1 and TCP21/CHE to repress their transcription. Represses both CCA1 and itself. {ECO:0000269|PubMed:12015970, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:9657154}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
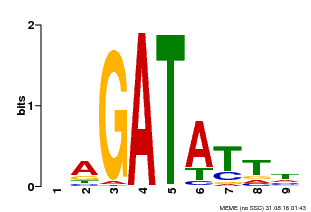
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation with peak levels occurring around 1 hour after dawn. Up-regulated by APRR1/TOC1 and transiently by light treatment. Down-regulated by APRR5, APRR7 and APRR9. {ECO:0000269|PubMed:12574129, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:19286557, ECO:0000269|PubMed:20233950, ECO:0000269|PubMed:9657154}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK242771 | 0.0 | AK242771.1 Oryza sativa Japonica Group cDNA, clone: J090053J23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015649844.1 | 0.0 | protein LHY isoform X2 | ||||
| Refseq | XP_015649845.1 | 0.0 | protein LHY isoform X2 | ||||
| Swissprot | Q6R0H1 | 4e-44 | LHY_ARATH; Protein LHY | ||||
| TrEMBL | A0A0E0EI42 | 0.0 | A0A0E0EI42_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI08G03470.2 | 0.0 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 4e-40 | circadian clock associated 1 | ||||



