 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KN541440.1_FGP002 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1072aa MW: 119404 Da PI: 7.9315 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.2 | 0.00027 | 953 | 976 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
++C C++sFs++ +L H r
KN541440.1_FGP002 953 FPCDieGCDMSFSTQQDLLLHKRD 976
789999***************985 PP
| |||||||
| 2 | zf-C2H2 | 12.2 | 0.00054 | 1036 | 1062 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
KN541440.1_FGP002 1036 YECQepGCGQTFRFVSDFSRHKRKtgH 1062
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00558 | 3.7E-29 | 1 | 149 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 29.653 | 1 | 149 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 6.87E-22 | 7 | 146 | No hit | No description |
| Pfam | PF02373 | 1.8E-34 | 12 | 132 | IPR003347 | JmjC domain |
| SMART | SM00355 | 9.7 | 953 | 975 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 15 | 976 | 1000 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.907 | 976 | 1005 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 978 | 1000 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.16E-14 | 987 | 1042 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 4.0E-11 | 1003 | 1030 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 8.6E-4 | 1006 | 1030 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.655 | 1006 | 1035 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1008 | 1030 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.1E-10 | 1031 | 1059 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.18 | 1036 | 1062 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.926 | 1036 | 1067 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1038 | 1062 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1072 aa Download sequence Send to blast |
MARDVAGVTT PMLYVAMLYS WFAWHVEDHE LHSLNFLHFG KAKTWYGVPR DAMLAFEETV 60 RVHGYADDLN AIRVAFQTLN EKTTVLSPEV LLSAGVPCCR LVQKAGEFVI TFPGAYHSGF 120 SHGFNCGEAS NIATPHWLQV AKEAAIRRAS TNCGPMVSHY QLLYELALSL RPREPKNFYS 180 VPRSSRLRDK NKNEGDIMVK ENFVGSVTEN NNLLSALLDK NSCIIVPNAD FFVPSFPVAL 240 ESEVTVKQRF TAGPCSISQQ GAENMAADHV AVDKVTEIQD MSGSLYPCET SLVGCSNRKL 300 YETKYGQRDA AALCLSTSEI QSRGIDTARS HPAGGILDQG RLPCVQCGIL SFACVAIIQP 360 REAAVQFIMS KECISSSAKQ GGIGASDDTS NWIDQSHEIS PPPGPASGTD DNVKHAVSLA 420 HVSDRCRELY ASNTDGCTSA LGLLASAYDS SDSDDETTED VSKHSKKNDS VNQSTDPQIL 480 ETSASCSSTV QCQKTNSHLH EEECEARATS LMKPLQQIGG ADIMLLCHPE YPRAESAAKV 540 IAEELGIKHD WKDITFKEAT EEDVKKIQLA LQDEDAEPTG SDWAVKMGIN IYYSAKQSKS 600 PLYSKQIPYN SIIYKAFGQE NPDSLTDYGC QKSGSTKKKV AGWWCGKVWM SNQVHPLLAR 660 EREEQNSSVV YGKAMFTTIS HGKVQDEAST RCNTSNRTPS RRTSRRKKGV SAEKSKPKNK 720 RSTASDEASM HCSGLGMNSG VIHDQTENSD DYDKHGNGDE IEEGINPQKY QQRKLQNVTR 780 KSSSKKRKDE KRTDSFHELY DEDNGVDYWL NMGSGDDATL GNSRQQSPDP VKVKSGGKLQ 840 GKRKSSKYKS NDDLLNEENK LQKMNKKSSS KKQKNDKINR QLQEDQTEDD HMDHLVDVAV 900 ADEVTLDNED KITEDKIDDV KVKSRGKSQN GKRKGSKHQA TDGLRAGNKV AKFPCDIEGC 960 DMSFSTQQDL LLHKRDICPV KGCKKKFFCH KYLLQHRKVH IDERPLKCTW KGCKKAFKWP 1020 WARTEHMRVH TGVRPYECQE PGCGQTFRFV SDFSRHKRKT GHSSDKRRKN ST |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 8e-57 | 950 | 1069 | 20 | 139 | Lysine-specific demethylase REF6 |
| 6a58_A | 8e-57 | 950 | 1069 | 20 | 139 | Lysine-specific demethylase REF6 |
| 6a59_A | 8e-57 | 950 | 1069 | 20 | 139 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
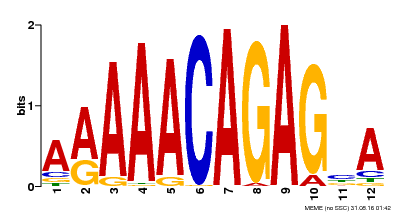
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KN541440.1_FGP002 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK065251 | 0.0 | AK065251.1 Oryza sativa Japonica Group cDNA clone:J013002J08, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015620626.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 1e-166 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A0E0BRB6 | 0.0 | A0A0E0BRB6_9ORYZ; Uncharacterized protein | ||||
| STRING | OGLUM12G09840.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6718 | 30 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-109 | relative of early flowering 6 | ||||



