 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KN538892.1_FGP004 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1031aa MW: 115266 Da PI: 6.3767 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 177.9 | 1.3e-55 | 22 | 137 | 3 | 118 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptf 94
+ +krwl+++ei++iL n+++++l+ e+++rp sgsl+L++rk +ryfrkDG++w+kkkdgktv+E+hekLK g+++vl+cyYah+een++f
KN538892.1_FGP004 22 EARKRWLRPTEICEILSNYRSFSLSPEPPNRPGSGSLFLFDRKTLRYFRKDGHNWRKKKDGKTVKEAHEKLKAGSIDVLHCYYAHGEENENF 113
459***************************************************************************************** PP
CG-1 95 qrrcywlLeeelekivlvhylevk 118
qrr+ywlLee++++ivlvhylevk
KN538892.1_FGP004 114 QRRTYWLLEEDFTHIVLVHYLEVK 137
*********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.074 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.9E-75 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.5E-49 | 22 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:1.25.40.20 | 3.2E-17 | 89 | 109 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00102 | 0.00522 | 449 | 537 | No hit | No description |
| Gene3D | G3DSA:2.60.40.10 | 2.3E-7 | 450 | 526 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.2E-7 | 450 | 535 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 3.89E-17 | 450 | 536 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 2.02E-16 | 626 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.45E-12 | 641 | 739 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.2E-17 | 641 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 18.033 | 647 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 4.0E-7 | 653 | 742 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.778 | 680 | 712 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.061 | 680 | 709 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 500 | 719 | 748 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.04E-8 | 847 | 905 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.0082 | 854 | 876 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.791 | 855 | 884 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0055 | 856 | 875 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0051 | 877 | 899 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.267 | 878 | 902 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 3.1E-4 | 880 | 899 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1031 aa Download sequence Send to blast |
MAETRKFLMP NQPPDISQMV LEARKRWLRP TEICEILSNY RSFSLSPEPP NRPGSGSLFL 60 FDRKTLRYFR KDGHNWRKKK DGKTVKEAHE KLKAGSIDVL HCYYAHGEEN ENFQRRTYWL 120 LEEDFTHIVL VHYLEVKGVK QSFSRSKEEI MQLSGADSPS CSNSITSQNQ MTPQIMDAAE 180 SPISGQISEY EGAEPAKFGA ADNCRASSRY NPLIEMQQPL DGIVMDSILY PSASAICNQV 240 SGYHGELPPG TSNLNGHTFT HSDIARMFDD SSSGLRDISR TLFDSMPYDE HLSGYANGFM 300 EPTLHSSFSM IEANNLEDSS LLETFTSEAL YTNNLSQKEA DALSFAGISS PEVNGNKYTE 360 GSTKHPLLKQ LPLDLFKIES SGLKKHDSFS RWMSKELGEV VDLGIKSSSD ALWSSTENVN 420 AADGPSAPIN EQLDAYAVSP SLAQDQLFSI LDISPSCSYI GLKTKVLVTG TFLASKENVE 480 NCKWSCMFGD VEVPAEVLAD GSLRCYAPEH QSGRVPFYVT CSNRIACSEV REFEYRDSDA 540 QYMETSHSQA NGINEMHLQI RLEKLLSLGP DDNQLLVCGN EKLELINAIN SLMLDEKWSD 600 QGSPSGSKDV VTPRNQSLKK LMKEKLHCWL IYKINDCEKG PNILGKEGQG IIHLVAALGF 660 DWAIRPILVA GVNVNFRDAH GWTALHWAAS CGRERTVGVL IANGAAAGAL TDPTSEFPSG 720 RTPADLASTN GHKGIAGFLA ESALTSHLSA LSLKDSKDSN VEEARGLTIP EDIPEMYSGQ 780 LAVQDSHAES LKDSLSAVRK SAQAAARIFQ AFRVESFHRK KVVEYGDDDC GLSDEHTFSL 840 ISLQKVKQGQ HDTRLHSAAV RIQNKFRGWK GRKEFMIIRQ RIVKLQAHVR GHQVRKNYKK 900 VVWSVGIVEK VILRWRRKGR GLRGFRPEKQ LEGQTQIQPA KTEDEYDYLQ DGRRQAEGRL 960 QRALDRVRSM TQYPEAREQY RRLTTCVAEM QQSRMMQDEM LSEAAGADGS DFMNGLEDLI 1020 CRDDPQMSAI W |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
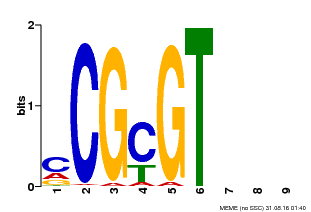
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KN538892.1_FGP004 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK070350 | 0.0 | AK070350.1 Oryza sativa Japonica Group cDNA clone:J023052K16, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015631089.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X1 | ||||
| TrEMBL | A0A0D9Z8A4 | 0.0 | A0A0D9Z8A4_9ORYZ; Uncharacterized protein | ||||
| STRING | OGLUM03G20490.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



