 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KN538684.1_FGP061 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 891aa MW: 100620 Da PI: 7.987 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.4 | 1.3e-05 | 582 | 604 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cg sF + +Lk+H+++H
KN538684.1_FGP061 582 HTCQECGASFQKPAHLKQHMQSH 604
79*******************99 PP
| |||||||
| 2 | zf-C2H2 | 20.6 | 1.2e-06 | 610 | 634 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s+ rk++L+rH+ +H
KN538684.1_FGP061 610 FICPleDCPFSYIRKDHLNRHMLKH 634
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 14.1 | 0.00013 | 639 | 664 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C+ Cg+ Fs k n++rH + H
KN538684.1_FGP061 639 FTCSmdGCGRKFSIKANMQRHVKEiH 664
89********************9988 PP
| |||||||
| 4 | zf-C2H2 | 14.4 | 0.00011 | 676 | 700 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ C+k+F+ s +k+H +H
KN538684.1_FGP061 676 FVCKeeGCNKVFKYASKMKKHEESH 700
89*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 16.5 | 2.5e-05 | 767 | 791 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C sFs+ksnL++Hi+ H
KN538684.1_FGP061 767 KCSfeGCECSFSNKSNLTKHIKAsH 791
688889***************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51375 | 8.703 | 91 | 125 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 2.7E-8 | 95 | 125 | IPR011990 | Tetratricopeptide-like helical domain |
| Gene3D | G3DSA:1.25.40.10 | 2.7E-8 | 160 | 207 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 9.701 | 161 | 195 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 1.2E-4 | 163 | 190 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 1.0E-4 | 163 | 191 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.086 | 194 | 221 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 10.063 | 223 | 257 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 8.3E-5 | 225 | 254 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 1.5E-4 | 225 | 258 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.903 | 293 | 323 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 2.7E-8 | 296 | 355 | IPR011990 | Tetratricopeptide-like helical domain |
| Pfam | PF01535 | 0.03 | 298 | 322 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF13041 | 1.3E-10 | 323 | 371 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 11.641 | 324 | 358 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 4.8E-6 | 326 | 359 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 9.219 | 359 | 389 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 0.001 | 361 | 388 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.774 | 395 | 425 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.044 | 398 | 422 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 2.7E-8 | 425 | 491 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 5.579 | 427 | 457 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.87 | 461 | 495 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.56 | 470 | 492 | IPR002885 | Pentatricopeptide repeat |
| SMART | SM00355 | 24 | 536 | 558 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-5 | 579 | 601 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0017 | 582 | 604 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.547 | 582 | 609 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 584 | 604 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 8.94E-11 | 594 | 636 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.6E-11 | 602 | 636 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 8.808 | 610 | 639 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.5E-4 | 610 | 634 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 612 | 634 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.92E-8 | 620 | 668 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-7 | 637 | 666 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 12.341 | 639 | 669 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.9E-4 | 639 | 664 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 641 | 664 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-7 | 673 | 700 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.764 | 676 | 700 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0038 | 676 | 700 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 678 | 700 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.7 | 708 | 733 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 710 | 733 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 736 | 757 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-5 | 748 | 788 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.19E-7 | 749 | 803 | No hit | No description |
| PROSITE profile | PS50157 | 11.365 | 766 | 796 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.002 | 766 | 791 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 768 | 791 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.8 | 797 | 823 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 799 | 823 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 891 aa Download sequence Send to blast |
MPKTDAYLGV RSRALRRASC VEERGMNRSV CHHLLTQCKT IRELQRIHAQ ALTHGLHPNQ 60 QSISCKIFRS YAEFGRPADA GRLFDEIPHP DIISFTSLMS LHLKLDHHRK AISVFSHAIA 120 SGHRPDGFAA VGALSASGGL GDHRIGSAVH GLIFRCGLDS ELVVCNALVD MYCRCGKFEP 180 ARTVFDRMLV KDEVTWGSML YGYMKCVGVD SALSFFYQMP MKSTVSWTAL ITGHVQDKQP 240 IQALELFGKM LLEGHRPTHI TIVGVLSACA DIGALDLGRA IHGYGSKSNA TTNIIVTNAL 300 MDMYAKSGSI ASAFSVFEEV QMKDAFTWTT MISSFTVQGN GRKAVELFWD MLRSGILPNS 360 VTFVSVLSAC SHAGLIQEGR ELFDKMREVY HIDPRLEHYG CMVDLLGRGG LLEEAEALID 420 HMDVEPDIVI WRSLLSACLA HGNDRLAEIA GKEIIKREPG DDGVYVLLWN MYASSNRWKE 480 ALDMRKQMLS KKIYKKPGCS WIEVDGVVHE FLMCSGDDID GDTRVEATQH RDIRRYKCEF 540 CTVVRSKKCL IRAHMVAHHK EELDKSEIYK SNGEKVVHEG DHTCQECGAS FQKPAHLKQH 600 MQSHSDERSF ICPLEDCPFS YIRKDHLNRH MLKHQGKLFT CSMDGCGRKF SIKANMQRHV 660 KEIHEDENAT KSNRQFVCKE EGCNKVFKYA SKMKKHEESH VKLDYVEVVC CEPGCMKTFT 720 NVECLRAHNQ ACHQYVQCDI CGEKHLKKNI KRHLRAHEEV PSTERIKCSF EGCECSFSNK 780 SNLTKHIKAS HDQVKPFACR FTGCEKVFPY KHVRDNHEKS SAHVYTQANF TEMDEQLLSC 840 PRGGRKRKAV TVETLTRKRV TMHGDASSLD NGTEYLRWLL SGGDDDSSQT H |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5iww_D | 7e-57 | 45 | 493 | 10 | 333 | PLS9-PPR |
| Search in ModeBase | ||||||
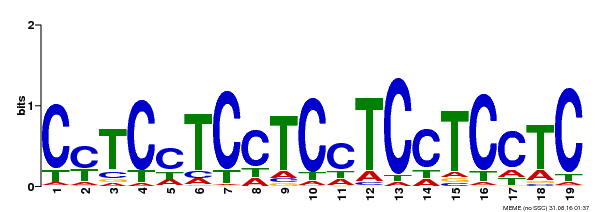
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KN538684.1_FGP061 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012610 | 0.0 | CP012610.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 2 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025878917.1 | 0.0 | pentatricopeptide repeat-containing protein At2g22410, mitochondrial | ||||
| TrEMBL | A0A0D3EZT2 | 0.0 | A0A0D3EZT2_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART02G01180.2 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-117 | transcription factor IIIA | ||||



