 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OGLUM09G03800.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | B3 | ||||||||
| Protein Properties | Length: 1523aa MW: 173751 Da PI: 9.3492 | ||||||||
| Description | B3 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 35.9 | 1.3e-11 | 108 | 184 | 16 | 98 |
EE--HHH.HTT.---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE- CS
B3 16 lvlpkkfaeeh.ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfr 98
++ k fa++ g s +le ++g++++v++ +k ++ v+++GW+ F+ a +L++gD++ F ++g+s f +v++f+
OGLUM09G03800.1 108 KTILKMFAKNVqG-L--ISGVAKLEVPDGKTYDVEI--SKEHNELVFRSGWEVFAIAYELEQGDILDFGYSGNSHF--KVQIFN 184
4788999977744.4..35579**************..**********************************9999..999986 PP
| |||||||
| 2 | B3 | 54.8 | 1.7e-17 | 229 | 320 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
ffkv+++ + k l +pkkfa++ gk + +++le ++g+ ++v++ + +++ vl++GW +F + +Lk gD +vF ++g+s f +v
OGLUM09G03800.1 229 FFKVMMSVSDVKDE-LAIPKKFAANVRGK--IPEQVRLEVSDGKMYNVQV--TEEQDELVLRSGWANFSSTYQLKHGDLLVFIHSGHSHF--KV 315
89***999999987.********888666..3446***************..********************************999999..99 PP
EEE-S CS
B3 95 kvfrk 99
+f+
OGLUM09G03800.1 316 LIFDP 320
99885 PP
| |||||||
| 3 | B3 | 36.3 | 1e-11 | 444 | 526 | 9 | 92 |
HHHHTT-EE--HHH.HTT---..--SEEEEEETTS.-EEEEEE..EEETTE.EEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE.. CS
B3 9 dvlksgrlvlpkkfaeehggkkeesktltledesg.rsWevkliyrkksgr.yvltkGWkeFvkangLkegDfvvFkldgr.sefel 92
+ +sg lv+ k++a ++ ++ +++t++l ++ g ++W++ l + + y l++GW +F + n L+egD++vF+ +++ ++ l
OGLUM09G03800.1 444 TSVASGSLVFCKDYAVRYLLD--QNRTIKLCQSGGsKTWDISL--DMDTDDlYALSTGWLDFFRGNLLQEGDICVFEASKSkRGVAL 526
4556778***********888..4558888887777*******..8888888*************************8876444444 PP
| |||||||
| 4 | B3 | 39.2 | 1.2e-12 | 588 | 686 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefel 92
f++v+ +s+ ++++ l+++ k+a+++ k + ++l+ e+ + + l ++ + + + l kGW +Fv++n+L+ D+++F+l+++ ++ +
OGLUM09G03800.1 588 FLSVMRSSNCTRQSSLCFSVKYASKYLPHKDQ--NMRLRLpETKYKCKAALHIDTSTNLHKLLKGWGKFVNDNKLEIHDICLFQLMKNkKKLTV 679
7889999999999*************544345..5666663455778888877999999***************************99999999 PP
EEEEE-S CS
B3 93 vvkvfrk 99
+v+++rk
OGLUM09G03800.1 680 TVHIIRK 686
9999987 PP
| |||||||
| 5 | B3 | 60.4 | 3.1e-19 | 715 | 802 | 1 | 97 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
f+kv+ +k+g +++p kfa+++gg+ s t++le+++g+++ev++ k + vl++GW++F++a +L+ gD++vF ++g+s f +v
OGLUM09G03800.1 715 FIKVM--ITDFKNG-VTIPAKFARNFGGQM--SGTVKLETRNGKTYEVQV--AKELNNLVLRSGWERFASAYELEKGDILVFIYSGNSHF--KV 799
67777..6678887.********9999994..559***************..**********************************9999..66 PP
EEE CS
B3 95 kvf 97
++
OGLUM09G03800.1 800 WIY 802
655 PP
| |||||||
| 6 | B3 | 40.5 | 4.8e-13 | 931 | 1012 | 8 | 93 |
-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTE.EEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..E CS
B3 8 sdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgr.yvltkGWkeFvkangLkegDfvvFkldgrsefelv 93
+ + + g+l ++k++a ++ ++k+e t++l + ++ +W++ l + + y l++GW +F + n+L+egD++vF+ ++++++ v
OGLUM09G03800.1 931 KASATDGLLAISKDYAVSYLLDKNE--TIKLCHsGRSMTWDISL--DIDTDDqYALSTGWLDFIRNNHLQEGDICVFEASKNKRG--V 1012
556677999**********888767..56665536678899999..4444434*************************8887666..3 PP
| |||||||
| 7 | B3 | 34.9 | 2.7e-11 | 1089 | 1169 | 13 | 96 |
TT-EE--HHH.HTT.---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE..EEEE CS
B3 13 sgrlvlpkkfaeeh.ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefelvvkv 96
s+ ++++ k+a+++ e++ +++ + ++s++ k ++ + + + l++GW +Fv++n++k gD+++F+l+++ ++ l+++v
OGLUM09G03800.1 1089 SRYMCFSAKYAAKYlP--REKN-KIMRLRLPNKSYRYKAVFKINNKVHKLGGGWGKFVDDNKIKLGDICLFQLMKNkKK--LMMMV 1169
5667888899999942..3344.455555566777777777999999**************************997444..44444 PP
| |||||||
| 8 | B3 | 44.5 | 2.9e-14 | 1207 | 1283 | 14 | 94 |
T-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 14 grlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
+ +++p k ae ++g+ ++ l+++sg++W+v + k ++ +l++GW++F+ka++L+e+D + F+ +g+ ++++ +
OGLUM09G03800.1 1207 HGISIPEKVAEIFSGQITK--GFNLKSPSGETWRVGV--AKVADELILKSGWEDFAKAHELQENDLLFFTCNGHGNGSCSF 1283
459*******777777555..59999***********..*******************************77665554443 PP
| |||||||
| 9 | B3 | 43.3 | 6.6e-14 | 1416 | 1511 | 1 | 96 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE...EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliy.rkksgryvltkGWkeFvkangLkegDfvvFkldgr.se 89
f+ vl +++v ++++l++p +fa++h + +s ++ l + ++W+vk+ y +++ +++ W +F ++n+L+eg +++F+l++ ++
OGLUM09G03800.1 1416 FVVVLQTAHVRRRNILIVPTRFAADHLER--KSHDILLIRpNRKQKWSVKYYYlSNTTRGFNCH-RWIKFIRENRLREGNVCIFELMKGaRR 1504
788999********************544..45567776636779******8865555555544.8*******************9975455 PP
E..EEEE CS
B3 90 felvvkv 96
++v+v
OGLUM09G03800.1 1505 PIMTVHV 1511
5556655 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 2.7E-14 | 94 | 186 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 0.0016 | 97 | 186 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 1.45E-14 | 98 | 185 | IPR015300 | DNA-binding pseudobarrel domain |
| Pfam | PF02362 | 1.4E-9 | 109 | 184 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 5.66E-13 | 112 | 184 | No hit | No description |
| PROSITE profile | PS50863 | 10.01 | 128 | 186 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 6.47E-21 | 222 | 322 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.6E-21 | 224 | 322 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.15E-17 | 227 | 319 | No hit | No description |
| PROSITE profile | PS50863 | 12.717 | 228 | 321 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 7.3E-10 | 229 | 321 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.1E-14 | 229 | 319 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 2.3E-14 | 429 | 533 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 5.49E-15 | 430 | 534 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.55E-10 | 435 | 532 | No hit | No description |
| PROSITE profile | PS50863 | 10.87 | 436 | 534 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.8E-4 | 438 | 534 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.4E-9 | 444 | 526 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 1.26E-17 | 581 | 686 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.0E-15 | 581 | 685 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.50E-14 | 586 | 682 | No hit | No description |
| SMART | SM01019 | 7.7E-7 | 588 | 687 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 9.164 | 588 | 687 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.0E-11 | 588 | 686 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 3.53E-22 | 709 | 807 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.3E-22 | 711 | 807 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 13.465 | 712 | 805 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 4.47E-20 | 713 | 803 | No hit | No description |
| SMART | SM01019 | 2.6E-16 | 715 | 805 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.1E-17 | 715 | 801 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 6.08E-17 | 918 | 1017 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.7E-16 | 918 | 1016 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.39E-10 | 923 | 1004 | No hit | No description |
| PROSITE profile | PS50863 | 12.012 | 924 | 1019 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 5.5E-6 | 925 | 1022 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.8E-10 | 933 | 1011 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 8.0E-15 | 1065 | 1177 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.29E-17 | 1070 | 1176 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.47E-13 | 1074 | 1156 | No hit | No description |
| SMART | SM01019 | 0.0013 | 1076 | 1173 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 9.7 | 1077 | 1173 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.3E-9 | 1088 | 1169 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 3.92E-17 | 1190 | 1293 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.069 | 1194 | 1291 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 1.66E-13 | 1196 | 1289 | No hit | No description |
| SMART | SM01019 | 1.4E-6 | 1197 | 1291 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 6.9E-17 | 1197 | 1293 | IPR015300 | DNA-binding pseudobarrel domain |
| Pfam | PF02362 | 1.4E-11 | 1206 | 1281 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 8.9E-17 | 1409 | 1513 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 9.42E-20 | 1410 | 1508 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.60E-17 | 1414 | 1510 | No hit | No description |
| SMART | SM01019 | 5.9E-9 | 1416 | 1513 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.014 | 1416 | 1515 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.4E-11 | 1416 | 1511 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005773 | Cellular Component | vacuole | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1523 aa Download sequence Send to blast |
MMNNTPCGHE GSMSYHDNHL QSPSESSSLF NISLPPHSPF QELSGVDSTS LLVSDPPNMQ 60 QFCLRCSWTN PKRLAKPSLA IASLSHQHLA FDKTRCMFIL KIENDTLKTI LKMFAKNVQG 120 LISGVAKLEV PDGKTYDVEI SKEHNELVFR SGWEVFAIAY ELEQGDILDF GYSGNSHFKV 180 QIFNPRASLM ITIDNHPEGK VWMNKPSTTC MDCITNHYWL HMDDRERYFF KVMMSVSDVK 240 DELAIPKKFA ANVRGKIPEQ VRLEVSDGKM YNVQVTEEQD ELVLRSGWAN FSSTYQLKHG 300 DLLVFIHSGH SHFKVLIFDP SCTEKEFSCV VTDNTSHVHE RSISHDNHLQ SPRSEILGKN 360 YSLCSSRKRS RMNPADYPSQ RPDVPSSEDI KDPMSSGGLQ KSKKSCYVLP MLYNMTSAQE 420 AEVLALEKKI QPQIPLYITA MDKTSVASGS LVFCKDYAVR YLLDQNRTIK LCQSGGSKTW 480 DISLDMDTDD LYALSTGWLD FFRGNLLQEG DICVFEASKS KRGVALTFHP FKESHCPKSS 540 EYTLSTKSPT RRVPKRDYFA TNLTNLTDQL ERKVKNKIKF IQSDIPIFLS VMRSSNCTRQ 600 SSLCFSVKYA SKYLPHKDQN MRLRLPETKY KCKAALHIDT STNLHKLLKG WGKFVNDNKL 660 EIHDICLFQL MKNKKKLTVT VHIIRKGEME KSHRVCKNCV ANHYWLHMDN HGKSFIKVMI 720 TDFKNGVTIP AKFARNFGGQ MSGTVKLETR NGKTYEVQVA KELNNLVLRS GWERFASAYE 780 LEKGDILVFI YSGNSHFKVW IYDPSACEKE LPCIITEQLP RVQQRSISHN NHTQLKRNAK 840 SAKLYVDSSG HSKETSEINP ANSPSWKPTE RVPSSEELDE PVDLANVQKA TKSFYSLPRM 900 CNMTSAQKAE VDALEKRIKP QIPLYITVMD KASATDGLLA ISKDYAVSYL LDKNETIKLC 960 HSGRSMTWDI SLDIDTDDQY ALSTGWLDFI RNNHLQEGDI CVFEASKNKR GVALIFHPLK 1020 QSHHPKPPGC VPSTKFPRHG VSKPNYIVSR FTTLSGQLKI KVEAKVQAMQ SEIPIFVAVM 1080 RESFIRGRSR YMCFSAKYAA KYLPREKNKI MRLRLPNKSY RYKAVFKINN KVHKLGGGWG 1140 KFVDDNKIKL GDICLFQLMK NKKKLMMMVM GEKGCESCRK WQEHYYREHM DVSRIRFFRL 1200 MTGDFAHGIS IPEKVAEIFS GQITKGFNLK SPSGETWRVG VAKVADELIL KSGWEDFAKA 1260 HELQENDLLF FTCNGHGNGS CSFDVLIFDA SGCEKVSCFF TGKKNSYMCK NFNSIGGQVA 1320 GQYLSSDSED TSTPSVLIGS PHKASTSKKL SGKTKTNPRK EPEDPNCSHW HVIEEKNTDD 1380 DEHADYHYTR FANYLTGEER DEIFSLVSLQ PGNPVFVVVL QTAHVRRRNI LIVPTRFAAD 1440 HLERKSHDIL LIRPNRKQKW SVKYYYLSNT TRGFNCHRWI KFIRENRLRE GNVCIFELMK 1500 GARRPIMTVH VIGKADNRFV LLG |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00468 | DAP | Transfer from AT4G33280 | Download |
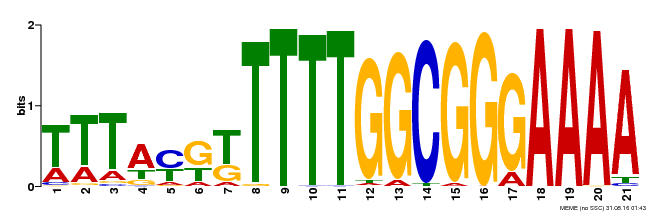
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK241993 | 0.0 | AK241993.1 Oryza sativa Japonica Group cDNA, clone: J075099J10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025878051.1 | 0.0 | B3 domain-containing protein LOC_Os12g40080 | ||||
| Swissprot | Q2QMT6 | 0.0 | Y1208_ORYSJ; B3 domain-containing protein LOC_Os12g40080 | ||||
| TrEMBL | A0A0E0B0J6 | 0.0 | A0A0E0B0J6_9ORYZ; Uncharacterized protein | ||||
| STRING | OGLUM09G03800.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP562 | 34 | 179 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G33280.1 | 7e-28 | B3 family protein | ||||



