 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OGLUM06G24700.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 426aa MW: 44261.9 Da PI: 9.3015 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.1 | 4e-41 | 180 | 256 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq+egC+adls ak+yhrrhkvCe h+ka+vv++sg++qrfCqqCsrfh l+efDe+krsCr+rLa+hn+rrrk++
OGLUM06G24700.1 180 RCQAEGCKADLSGAKHYHRRHKVCEYHAKASVVAASGKQQRFCQQCSRFHVLTEFDEAKRSCRKRLAEHNRRRRKPA 256
6**************************************************************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.6E-35 | 173 | 242 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.547 | 178 | 255 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.06E-38 | 179 | 257 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.2E-31 | 181 | 254 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 426 aa Download sequence Send to blast |
MMSGRMNAAG DESPFPFGAM QAPGPGAYVG FDHGAAAVAA AAAAAQRAGM LQHHHHHMYD 60 GLDFAAAMQF GGGQDAPPHP QLLALPPSMA APPPPPMPMP LQMPMTMPMP GDVYPALGIV 120 KREGGGGGQD AAAGRIGLNL GRRTYFSPGD MLAVDRLLMR SRLGGVFGLG FGGAHHQPPR 180 CQAEGCKADL SGAKHYHRRH KVCEYHAKAS VVAASGKQQR FCQQCSRFHV LTEFDEAKRS 240 CRKRLAEHNR RRRKPAAAPT TAVAAAKDAV AAPVAAGKKP SGGAATSYTG DNKNVVSMSA 300 AKSPISSNTS VISCLAEQGK HAAAAARPTA LTLGGAPPHE SSAPQIGAML HHHHHHQQDH 360 MQVSSLVHIN GGGGGGGSNN ILSCSSVCSS ALPSTATNGE VSNQNNDNSH NNGGNNNMHL 420 FEVDFM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 5e-30 | 181 | 254 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 237 | 254 | KRSCRKRLAEHNRRRRKP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
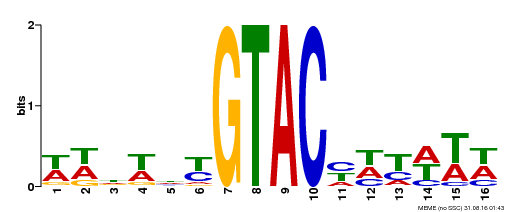
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP005760 | 0.0 | AP005760.3 Oryza sativa Japonica Group genomic DNA, chromosome 6, BAC clone:B1047G05. | |||
| GenBank | AP014962 | 0.0 | AP014962.1 Oryza sativa Japonica Group DNA, chromosome 6, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012614 | 0.0 | CP012614.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 6 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015642406.1 | 0.0 | squamosa promoter-binding-like protein 10 | ||||
| Swissprot | A2YFT9 | 0.0 | SPL10_ORYSI; Squamosa promoter-binding-like protein 10 | ||||
| Swissprot | Q0DAE8 | 0.0 | SPL10_ORYSJ; Squamosa promoter-binding-like protein 10 | ||||
| TrEMBL | A0A0E0ACS0 | 0.0 | A0A0E0ACS0_9ORYZ; Uncharacterized protein | ||||
| STRING | OGLUM06G24700.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2375 | 35 | 93 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 1e-45 | squamosa promoter binding protein-like 8 | ||||



