 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OGLUM05G13600.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 268aa MW: 27587.8 Da PI: 7.347 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.8 | 6.4e-19 | 49 | 98 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+WvAeIr+p++ r r +lg+f tae+Aa+a+++a++ l+g
OGLUM05G13600.1 49 RYRGVRQRS-WGKWVAEIREPRK---RSRKWLGTFATAEDAARAYDRAALLLYG 98
59****999.**********954...5**********************98877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00380 | 2.1E-36 | 49 | 112 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.2E-29 | 49 | 105 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.379 | 49 | 106 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.2E-12 | 49 | 97 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 5.26E-17 | 49 | 104 | No hit | No description |
| SuperFamily | SSF54171 | 7.19E-21 | 49 | 106 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 2.4E-10 | 50 | 61 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.4E-10 | 72 | 88 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006970 | Biological Process | response to osmotic stress | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0009749 | Biological Process | response to glucose | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010353 | Biological Process | response to trehalose | ||||
| GO:0010449 | Biological Process | root meristem growth | ||||
| GO:0010896 | Biological Process | regulation of triglyceride catabolic process | ||||
| GO:0031930 | Biological Process | mitochondria-nucleus signaling pathway | ||||
| GO:0032880 | Biological Process | regulation of protein localization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 268 aa Download sequence Send to blast |
MEPSDDACTV AAPAAETAAS SSGAGGGGGG GRTKKKAAGK GGPENGKFRY RGVRQRSWGK 60 WVAEIREPRK RSRKWLGTFA TAEDAARAYD RAALLLYGPR AHLNLTAPPP LPPPPPSSAA 120 AAAASSSSAA SSTSAPPPPL RPLLPRPPHL HPAFHHQPFH HHLLQPQPPP PLYYAATAST 180 STVTTTTTAP PPQLAAAAPA AVLVAAAVSS TAETQAVVAT APEDAASAAA AAAEEEEAAW 240 GFHGGDEEDY AAALLWSEPD PWFDLFLK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-16 | 50 | 105 | 6 | 62 | ATERF1 |
| 3gcc_A | 2e-16 | 50 | 105 | 6 | 62 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription regulator that binds to cis-responsive elements of genes involved in stress response. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00674 | SELEX | Transfer from GRMZM2G093595 | Download |
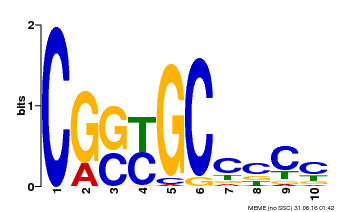
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC136221 | 0.0 | AC136221.2 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone OSJNBa0077J17, complete sequence. | |||
| GenBank | AP014961 | 0.0 | AP014961.1 Oryza sativa Japonica Group DNA, chromosome 5, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025881609.1 | 2e-66 | ethylene-responsive transcription factor ABI4 | ||||
| Swissprot | C7J2Z1 | 2e-67 | ABI4_ORYSJ; Ethylene-responsive transcription factor ABI4 | ||||
| TrEMBL | A0A0D9ZXV1 | 0.0 | A0A0D9ZXV1_9ORYZ; Uncharacterized protein | ||||
| STRING | OGLUM05G13600.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP13303 | 26 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40220.1 | 1e-18 | ERF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



