 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB02G35240.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 281aa MW: 31140.4 Da PI: 8.6986 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 42.8 | 1.2e-13 | 34 | 78 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a +++ + Wk+I + +g +t q++s+ qky
OB02G35240.1 34 RESWTEQEHDKFLEALQLFDRD-WKKIEAFVG-SKTVIQIRSHAQKY 78
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.22E-17 | 28 | 84 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.408 | 29 | 83 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.2E-18 | 32 | 81 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 8.8E-7 | 32 | 84 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.3E-11 | 33 | 81 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-11 | 34 | 78 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.39E-8 | 36 | 79 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 281 aa Download sequence Send to blast |
MVSANQPPPD AAGAASGEDA SKKGRKPYTI TKSRESWTEQ EHDKFLEALQ LFDRDWKKIE 60 AFVGSKTVIQ IRSHAQKYFL KVQKNGTSEH VPPPRPKRKA AHPYPQKASK NEPGYTLKTE 120 SSSMLRNSGM NATVSSWTHN SIPPIVASSM VKDLGAGAMG PNNFCSSSTE GPPRAWQPGE 180 TNDQINQVLS LRLMPDFAQV YSFLGSVFDP STSGHLQKLK EMNPIDVETA LLLMRNLSIN 240 LTSPDFEDQG LLICHGFWYC LCRMAGMYGQ LSENVCKVTI F |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00625 | PBM | Transfer from PK02532.1 | Download |
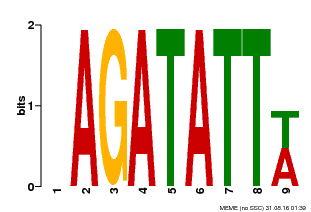
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB02G35240.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK288059 | 0.0 | AK288059.1 Oryza sativa Japonica Group cDNA, clone: J075153L04, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015689659.1 | 0.0 | PREDICTED: protein REVEILLE 6 isoform X2 | ||||
| Swissprot | Q8H0W3 | 1e-100 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | J3LFV7 | 0.0 | J3LFV7_ORYBR; Uncharacterized protein | ||||
| STRING | OB02G35240.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1096 | 38 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.2 | 1e-85 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



