 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART12G17530.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | B3 | ||||||||
| Protein Properties | Length: 1535aa MW: 175181 Da PI: 9.3526 | ||||||||
| Description | B3 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 38 | 2.9e-12 | 108 | 184 | 16 | 98 |
EE--HHH.HTT.---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE- CS
B3 16 lvlpkkfaeeh.ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfr 98
++ k fa++ g s +le ++g++++v++ +k ++ v+++GW+ F+ a +L++gD++ F ++g+s f +v++f+
OBART12G17530.1 108 KTILKMFAKNVqG-L--ISGVAKLEVPDGKTYDVEI--SKEHNELVFRSGWEVFAIAYELEQGDILAFGYSGNSHF--KVQIFN 184
4788999977744.4..35579**************..**********************************9999..999987 PP
| |||||||
| 2 | B3 | 54.7 | 1.8e-17 | 241 | 332 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
ffkv+++ + k l +pkkfa++ gk + +++le ++g+ ++v++ + +++ vl++GW +F + +Lk gD +vF ++g+s f +v
OBART12G17530.1 241 FFKVMMSVSDIKDE-LAIPKKFAANVRGK--IPEQVRLEVSDGKMYNVQV--TEEQDELVLRSGWANFSSTYQLKHGDLLVFIHSGHSHF--KV 327
99***999999987.********888666..3446***************..********************************999999..99 PP
EEE-S CS
B3 95 kvfrk 99
+f+
OBART12G17530.1 328 LIFDP 332
99885 PP
| |||||||
| 3 | B3 | 35.6 | 1.7e-11 | 456 | 535 | 9 | 89 |
HHHHTT-EE--HHH.HTT---..--SEEEEEETTS.-EEEEEE..EEETTE.EEE-TTHHHHHHHHT--TT-EEEEEE-SS.SE CS
B3 9 dvlksgrlvlpkkfaeehggkkeesktltledesg.rsWevkliyrkksgr.yvltkGWkeFvkangLkegDfvvFkldgr.se 89
+ +sg lv++k++a ++ ++ +++t++l ++ g ++W++ l + + y l++GW +F + n L+egD++vF+ + + ++
OBART12G17530.1 456 TSVASGSLVFSKDYAVRYLLD--QNRTIKLCQSGGsKTWDISL--DMDTDDlYALSTGWLDFFRGNLLQEGDICVFEASMSkRG 535
4556778***********888..4558888887777*******..8888888*************************7644244 PP
| |||||||
| 4 | B3 | 40.3 | 5.8e-13 | 600 | 698 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE.. CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefel 92
f++v+ +s+ ++++ l+++ k+a+++ k + ++l+ e+ + + l ++ + + + l kGW +Fv++n+L+ D+++F+l+++ ++ ++
OBART12G17530.1 600 FLSVMRSSNCTRQSSLCFSVKYASKYLPHKDQ--NMRLRLpETKYKCKAALHIDTSTNLHKLLKGWGKFVNDNKLEIHDICLFQLMKNkKKLTM 691
7889999999999*************544345..5666663455778888877999999***************************999***** PP
EEEEE-S CS
B3 93 vvkvfrk 99
+v+++rk
OBART12G17530.1 692 TVHIIRK 698
****997 PP
| |||||||
| 5 | B3 | 60.4 | 3.1e-19 | 727 | 814 | 1 | 97 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
f+kv+ +k+g +++p kfa+++gg+ s t++le+++g+++ev++ k + vl++GW++F++a +L+ gD++vF ++g+s f +v
OBART12G17530.1 727 FIKVM--ITDFKNG-VTIPAKFARNFGGQM--SGTVKLETRNGKTYEVQV--AKELNNLVLRSGWERFASAYELEKGDILVFIYSGNSHF--KV 811
67777..6678887.********9999994..559***************..**********************************9999..66 PP
EEE CS
B3 95 kvf 97
++
OBART12G17530.1 812 WIY 814
655 PP
| |||||||
| 6 | B3 | 40.9 | 3.8e-13 | 941 | 1024 | 6 | 93 |
..-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTE.EEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..E CS
B3 6 tpsdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgr.yvltkGWkeFvkangLkegDfvvFkldgrsefelv 93
+ + + + g+l ++k++a ++ ++k+e t++l + ++ +W++ l + + y l++GW +F + n+L+egD++vF+ ++++++ v
OBART12G17530.1 941 MDKASATDGLLAISKDYAVSYLLDKNE--TIKLCHsGRSMTWDISL--DIDTDDqYALSTGWLDFIRNNHLQEGDICVFEASKNKRG--V 1024
5566778899***********888767..56665536678899999..4444434*************************8887666..3 PP
| |||||||
| 7 | B3 | 35.3 | 2e-11 | 1101 | 1181 | 13 | 96 |
TT-EE--HHH.HTT.---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE..EEEE CS
B3 13 sgrlvlpkkfaeeh.ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefelvvkv 96
s+ ++++ k+a+++ e++ +++ + ++s++ k ++ + + + l++GW +Fv++n++k gD+++F+l+++ ++ l+++v
OBART12G17530.1 1101 SRYMCFSAKYAAKYlP--REKN-KIMRLRLPNKSYKYKAVFKINNKVHKLGGGWGKFVDDNKIKLGDICLFQLMKNkKK--LMMMV 1181
5667888899999942..3344.455555566777777777999999**************************997444..44444 PP
| |||||||
| 8 | B3 | 44.5 | 2.9e-14 | 1219 | 1295 | 14 | 94 |
T-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 14 grlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
+ +++p k ae ++g+ ++ l+++sg++W+v + k ++ +l++GW++F+ka++L+e+D + F+ +g+ ++++ +
OBART12G17530.1 1219 HGISIPEKVAEIFSGQITK--GFNLKSPSGETWRVGV--AKVADELILKSGWEDFAKAHELQENDLLFFTCNGHGNGSCSF 1295
459*******777777555..59999***********..*******************************77665554443 PP
| |||||||
| 9 | B3 | 45.4 | 1.5e-14 | 1428 | 1525 | 1 | 98 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE...EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliy.rkksgryvltkGWkeFvkangLkegDfvvFkldgr.se 89
f+ vl +++v ++++l++p +fa++h + +s ++ l + ++W+vk+ y +++ +++ W +F ++n+L+eg +++F+l++ ++
OBART12G17530.1 1428 FVVVLQTAHVRRRNILIVPTRFAADHLER--KSHDILLIRpNRKQKWSVKYYYlSNTTRGFNCH-RWIKFIRENRLREGNVCIFELMKGaRR 1516
788999********************544..45567776636779******8865555555544.8*******************9977688 PP
E..EEEEE- CS
B3 90 felvvkvfr 98
+++v+v+
OBART12G17530.1 1517 PTMTVHVIG 1525
889998876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.8E-15 | 94 | 188 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 4.0E-5 | 97 | 186 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 5.49E-16 | 98 | 188 | IPR015300 | DNA-binding pseudobarrel domain |
| Pfam | PF02362 | 3.9E-10 | 109 | 184 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 6.34E-14 | 112 | 184 | No hit | No description |
| PROSITE profile | PS50863 | 10.278 | 128 | 186 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 6.87E-21 | 234 | 334 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 3.4E-21 | 236 | 334 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.08E-17 | 239 | 331 | No hit | No description |
| PROSITE profile | PS50863 | 12.675 | 240 | 333 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.9E-10 | 241 | 333 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.1E-14 | 241 | 331 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 5.0E-14 | 441 | 544 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 8.83E-15 | 442 | 546 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.21E-11 | 447 | 544 | No hit | No description |
| PROSITE profile | PS50863 | 11.251 | 448 | 546 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.3E-4 | 450 | 546 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 9.0E-9 | 456 | 536 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 1.6E-16 | 592 | 698 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.53E-18 | 593 | 699 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.94E-14 | 598 | 697 | No hit | No description |
| SMART | SM01019 | 3.3E-6 | 600 | 699 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.5E-12 | 600 | 698 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 9.248 | 600 | 699 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 3.53E-22 | 721 | 819 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.3E-22 | 723 | 819 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 13.465 | 724 | 817 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 3.73E-20 | 725 | 815 | No hit | No description |
| SMART | SM01019 | 2.6E-16 | 727 | 817 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.1E-17 | 727 | 813 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 1.2E-16 | 930 | 1028 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.84E-17 | 930 | 1029 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.66E-10 | 935 | 1016 | No hit | No description |
| PROSITE profile | PS50863 | 12.379 | 936 | 1031 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 5.5E-6 | 937 | 1034 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.7E-10 | 942 | 1023 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 3.5E-15 | 1080 | 1182 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 9.42E-18 | 1081 | 1186 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.61E-13 | 1086 | 1168 | No hit | No description |
| SMART | SM01019 | 5.3E-4 | 1088 | 1185 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 9.77 | 1089 | 1185 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.1E-9 | 1100 | 1181 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 3.92E-17 | 1202 | 1305 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.069 | 1206 | 1303 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 1.45E-13 | 1208 | 1301 | No hit | No description |
| Gene3D | G3DSA:2.40.330.10 | 7.0E-17 | 1209 | 1305 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 1.4E-6 | 1209 | 1303 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.4E-11 | 1218 | 1293 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 4.0E-17 | 1421 | 1525 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 6.47E-20 | 1422 | 1523 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.43E-17 | 1426 | 1522 | No hit | No description |
| SMART | SM01019 | 7.9E-10 | 1428 | 1527 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.4E-12 | 1428 | 1525 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.225 | 1428 | 1527 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005773 | Cellular Component | vacuole | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1535 aa Download sequence Send to blast |
MMNNTPCGHE GSMSYHDNHL QSPSESSLLF NISLPPHSPF QELSGVDSTS LLVSDPTNMQ 60 QFCLRCSWTN PKRLAKPSLA IASLSHQHLA FDKTRCMFIL KIENDTLKTI LKMFAKNVQG 120 LISGVAKLEV PDGKTYDVEI SKEHNELVFR SGWEVFAIAY ELEQGDILAF GYSGNSHFKV 180 QIFNPSNCEK ELSCVVMNRS ISDDNHRQSP RRERMNKPST TCMDCITNHY WLHMDDRERY 240 FFKVMMSVSD IKDELAIPKK FAANVRGKIP EQVRLEVSDG KMYNVQVTEE QDELVLRSGW 300 ANFSSTYQLK HGDLLVFIHS GHSHFKVLIF DPSCTEKEFS CVVTDNTSHV HERSISHDNH 360 LQSPRSEILG KNYSLCSSRK RSRMNPADYP SQRPDVPSSE DIKDPMSSGG LQKSKKSCYV 420 LPMLYNMTSA QEAEVLALEK KIQPQIPLYI TAMDKTSVAS GSLVFSKDYA VRYLLDQNRT 480 IKLCQSGGSK TWDISLDMDT DDLYALSTGW LDFFRGNLLQ EGDICVFEAS MSKRGVALTF 540 HPFKESHCPK SSEYTLSTKS PTRRVPKRDY FATNLTNLTD QQERKVKNKI KSIQSDIPIF 600 LSVMRSSNCT RQSSLCFSVK YASKYLPHKD QNMRLRLPET KYKCKAALHI DTSTNLHKLL 660 KGWGKFVNDN KLEIHDICLF QLMKNKKKLT MTVHIIRKGE MEKSHRVCKN CVANHYWLHM 720 DNHGKSFIKV MITDFKNGVT IPAKFARNFG GQMSGTVKLE TRNGKTYEVQ VAKELNNLVL 780 RSGWERFASA YELEKGDILV FIYSGNSHFK VWIYDPSACE KELPCIITEQ LPRVQQRSIS 840 HNNHTQLKRN AKSAKLYVDS SGHSKETSEI NPANSPSWKP TERVPSSEEL DEPVDLANVQ 900 KATKSFYSLP RMCNMTSAQK AEVDALEKRI KPQIPFYITV MDKASATDGL LAISKDYAVS 960 YLLDKNETIK LCHSGRSMTW DISLDIDTDD QYALSTGWLD FIRNNHLQEG DICVFEASKN 1020 KRGVALIFHP LKQSHHPKPP GCVPSTKFPR HGVSKPNYIV SRFTTLSGQL KIKVEAKVQA 1080 IQSEIPIFVA VMRESFIRGR SRYMCFSAKY AAKYLPREKN KIMRLRLPNK SYKYKAVFKI 1140 NNKVHKLGGG WGKFVDDNKI KLGDICLFQL MKNKKKLMMM VMGEKGCESC RKWQEHYYRE 1200 HMDVSRIRFF RLMTGDFAHG ISIPEKVAEI FSGQITKGFN LKSPSGETWR VGVAKVADEL 1260 ILKSGWEDFA KAHELQENDL LFFTCNGHGN GSCSFDVLIF DASGCEKVSC FFTGKKNSYM 1320 CKNFNSIGGQ VAGQYLSSDS EDTSTPSVLI GSPHKASTSK KLSGKTKTNP RKEPEDPNCS 1380 HWHVIEEKNT DDDEHADYHY TRFANYLTGE ERDEIFSLVS LQPGNPVFVV VLQTAHVRRR 1440 NILIVPTRFA ADHLERKSHD ILLIRPNRKQ KWSVKYYYLS NTTRGFNCHR WIKFIRENRL 1500 REGNVCIFEL MKGARRPTMT VHVIGKADNR FVLLG |
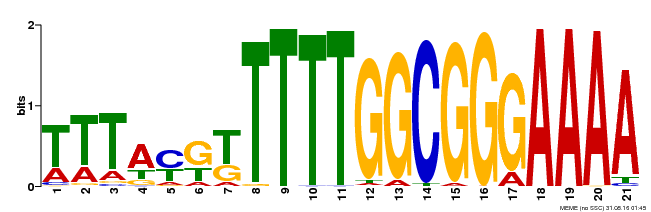
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00468 | DAP | Transfer from AT4G33280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK241993 | 0.0 | AK241993.1 Oryza sativa Japonica Group cDNA, clone: J075099J10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025878051.1 | 0.0 | B3 domain-containing protein LOC_Os12g40080 | ||||
| Swissprot | Q2QMT6 | 0.0 | Y1208_ORYSJ; B3 domain-containing protein LOC_Os12g40080 | ||||
| TrEMBL | A0A0D3HWA3 | 0.0 | A0A0D3HWA3_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART12G17530.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP562 | 34 | 179 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G33280.1 | 1e-27 | B3 family protein | ||||



