 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART12G08590.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1425aa MW: 158774 Da PI: 7.643 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.8 | 0.00037 | 1306 | 1329 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
++C C++sFs++ +L H r
OBART12G08590.1 1306 FPCDieGCDMSFSTQQDLLLHKRD 1329
789999***************985 PP
| |||||||
| 2 | zf-C2H2 | 11.8 | 0.00074 | 1389 | 1415 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
OBART12G08590.1 1389 YECQepGCGQTFRFVSDFSRHKRKtgH 1415
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.9E-16 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.201 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 3.6E-14 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 3.3E-43 | 241 | 411 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 32.594 | 241 | 411 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.57E-22 | 257 | 408 | No hit | No description |
| Pfam | PF02373 | 2.6E-34 | 274 | 394 | IPR003347 | JmjC domain |
| SMART | SM00355 | 9.7 | 1306 | 1328 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.907 | 1329 | 1358 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 1329 | 1353 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1331 | 1353 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.56E-14 | 1340 | 1394 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.5E-11 | 1356 | 1383 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 8.6E-4 | 1359 | 1383 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.655 | 1359 | 1388 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1361 | 1383 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.6E-10 | 1384 | 1412 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.18 | 1389 | 1415 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.926 | 1389 | 1420 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1391 | 1415 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1425 aa Download sequence Send to blast |
MSLQPPAVEP PEWLRTLPVA PEYHPTLAEF ADPIAYILRI EPEASRYGIC KIVPPLPRPP 60 EDDTFRRLQD AFAAAASSNG AGGDPSPTFP TRLQQVGLSA RNRRAASRRV WESGERYTLE 120 AFRAKAAEFE PPRHAAPPRN PTHLQLEALF WAACASRPFS VDRLVWESGE RYTLEAFRAK 180 AAEFEPSRHA TPPKNPTHLQ LEALFWAACT SRPFSVEYGN DMPGSAFASP DELPDAANAT 240 DVGETEWNMR VAPRARGSLL RAMARDVAGV TTPMLYVAML YSWFAWHVED HELHSLNFLH 300 FGKAKTWYGV PRDAMLAFEE TVRVHGYADD LNAIRVAFQT LNEKTTVLSP EVLLSAGVPC 360 CRLVQKAGEF VITFPGAYHS GFSHGFNCGE ASNIATPHWL QVAKEAAIRR ASTNCGPMVS 420 HYQLLYELAL SLRPREPKNF YSVPRSSRLR DKNKNEGDIM VKENFVGSVT ENNNLLSVLL 480 DKNYCIIVPN TDFFVPSFPV ALESEATVKQ RFTAGPCSIS QQGAENMAVD HVAVDKVTDI 540 QDMSGSLYPC ETSLVGCSNR KLYETKYGQR DAAALCLSTS EIQSRGIDKA RSHPAGGILD 600 QGRLPCVQCG ILSFACVAII QPREAAVQFI MSKECISSSA KQGGIGASDD TSNWIDQSHE 660 ISPPPGPASG TDDNVKHTVS LAHVSDRCRE LYASNTDGCT SALGLLASAY DSSDSDDETT 720 EDVLKHSKKN DSVNQSTDTQ ILETSASCSS TVRCQKTNSH SHEEECEARA TSLMKPVSHN 780 SRPISQSNRD TDIDQFIELG KSGTQCSGYL DLVDDLTTSV LKSSSDTCVS AAKASMDPDV 840 LTMLRYNKDS CRMHVFCLEH ALETWTQLQQ IGGANIMLLC HPEYPRAESA AKVIAEELGI 900 KHDWKDITFK EATEEDVKKI RLALQDEDAE PTGSDWAVKM GINIYYSAKQ SKSPLYSKQI 960 PYNSIIYKAF GQENPDSLTD YGCQKSGSTK KKVAGWWCGK VWMSNQVHPL LAREREEQNS 1020 SVVYGKAMFT TISHGKVQDE ASTRCNTSNR TPSKRTSRRK KGVSAEKSKP KNKRSTASDE 1080 VSMHCSGLGM NSGVIHDQTE NSDDYDKHGN GDEIEEGTNP QKYQQRKLQN VTRKSSSKKR 1140 KDEKRTDSFH ELYDEDNGVD YWLNMGSGHD ATLGNSQQQS PDPVKVKSGG KLQGKRKSSK 1200 YKSNDDLLNE ENKLQKMNKK SSSKKQKNDK INRQLQEDQT EDDHMDHLVD VAVADEVTLD 1260 NEDKITEDKI DDVKVKSRGK SQNGKRKGSK HQASDGLRTG NKVAKFPCDI EGCDMSFSTQ 1320 QDLLLHKRDI CPVKGCKKKF FCHKYLLQHR KVHIDERPLK CTWKGCKKAF KWPWARTEHM 1380 RVHTGVRPYE CQEPGCGQTF RFVSDFSRHK RKTGHSSDKR RKNST |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 1e-63 | 12 | 432 | 7 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 1e-63 | 12 | 432 | 7 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
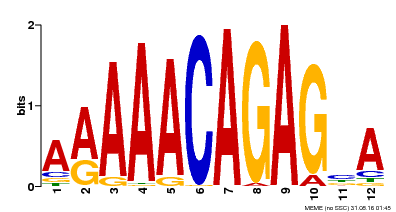
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK065251 | 0.0 | AK065251.1 Oryza sativa Japonica Group cDNA clone:J013002J08, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015620626.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A0D3HTA8 | 0.0 | A0A0D3HTA8_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART12G08590.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6718 | 30 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-151 | relative of early flowering 6 | ||||



