 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART09G12810.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 274aa MW: 29189.7 Da PI: 6.2795 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 63.3 | 5e-20 | 129 | 179 | 2 | 56 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkklege 56
y+GVr ++ +g+W+AeIrdp+ r r +lg+f+taeeAa+a+++a+++++g+
OBART09G12810.1 129 VYRGVRHRP-WGKWAAEIRDPR----RaVRKWLGTFDTAEEAARAYDRAALEFRGA 179
69****888.**********82....48*************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00380 | 3.5E-34 | 129 | 192 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 25.093 | 129 | 186 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.5E-13 | 129 | 178 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 7.19E-23 | 129 | 188 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 1.3E-31 | 129 | 187 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 7.65E-31 | 130 | 186 | No hit | No description |
| PRINTS | PR00367 | 5.3E-11 | 130 | 141 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.3E-11 | 152 | 168 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001944 | Biological Process | vasculature development | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 274 aa Download sequence Send to blast |
MTKKVIPAMA AARQDSCKTK LDERGGSHQA PSSARWISSE QEHSIIVAAL RYVVSGCTTP 60 PPEIVTVACG EACALCGIDG CLGCDFFGAE AAGNEEAVMA TDYAAAAAAA AVAGGSGGKR 120 VRRRRKKNVY RGVRHRPWGK WAAEIRDPRR AVRKWLGTFD TAEEAARAYD RAALEFRGAR 180 AKLNFPCSEP LPMPSQRNGN GGDAVTAATT TAEQMTPTLS PCSADAEETT TPVDWQMGAD 240 EAGSNQLWDG LQDLMKLDEA DTWFPPFSGA ASSF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-20 | 126 | 187 | 2 | 64 | ATERF1 |
| 3gcc_A | 1e-20 | 126 | 187 | 2 | 64 | ATERF1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 118 | 126 | KRVRRRRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
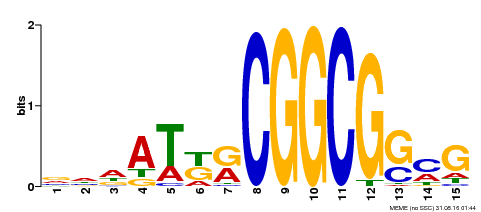
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00469 | DAP | Transfer from AT4G34410 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK067195 | 0.0 | AK067195.1 Oryza sativa Japonica Group cDNA clone:J013098P10, full insert sequence. | |||
| GenBank | AP005891 | 0.0 | AP005891.3 Oryza sativa Japonica Group genomic DNA, chromosome 9, BAC clone:B1045B05. | |||
| GenBank | AP014965 | 0.0 | AP014965.1 Oryza sativa Japonica Group DNA, chromosome 9, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012617 | 0.0 | CP012617.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 9 sequence. | |||
| GenBank | EU622934 | 0.0 | EU622934.1 Oryza sativa Indica Group EATB (EATB) mRNA, complete cds. | |||
| GenBank | KP212899 | 0.0 | KP212899.1 Oryza sativa Indica Group cultivar IR64-Sub1 AP2 domain containing protein (EATB) gene, complete cds. | |||
| GenBank | KP212900 | 0.0 | KP212900.1 Oryza sativa Indica Group cultivar PS-2 AP2 domain containing protein (EATB) gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015611044.2 | 0.0 | ethylene-responsive transcription factor ERF109 | ||||
| Swissprot | Q9SZ06 | 2e-30 | EF109_ARATH; Ethylene-responsive transcription factor ERF109 | ||||
| TrEMBL | A0A0D3H7P9 | 0.0 | A0A0D3H7P9_9ORYZ; Uncharacterized protein | ||||
| TrEMBL | A0A0E0QSC9 | 0.0 | A0A0E0QSC9_ORYRU; Uncharacterized protein | ||||
| TrEMBL | B2MV91 | 0.0 | B2MV91_ORYSI; AP2 domain containing protein | ||||
| TrEMBL | I1QPK1 | 0.0 | I1QPK1_ORYGL; Uncharacterized protein | ||||
| TrEMBL | Q67U00 | 0.0 | Q67U00_ORYSJ; Ethylene-binding protein-like | ||||
| STRING | ORUFI09G13770.1 | 0.0 | (Oryza rufipogon) | ||||
| STRING | OS09T0457900-01 | 0.0 | (Oryza sativa) | ||||
| STRING | ORGLA09G0098000.1 | 0.0 | (Oryza glaberrima) | ||||
| STRING | OBART09G12810.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1242 | 36 | 124 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34410.1 | 1e-21 | redox responsive transcription factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



