 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART07G22710.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1038aa MW: 115682 Da PI: 5.7341 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 180.4 | 2.2e-56 | 21 | 137 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
l+ ++rwl+++ei++iL n++k++++ e+++rp sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fq
OBART07G22710.1 21 LEAQNRWLRPTEICHILSNYKKFSIAPEPPNRPASGSLFLFDRKILRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQ 114
5679****************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+ywlLee + +ivlvhylevk
OBART07G22710.1 115 RRTYWLLEEGFMNIVLVHYLEVK 137
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.093 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.5E-76 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.4E-50 | 22 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 6.6E-5 | 442 | 529 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 1.71E-16 | 443 | 529 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 1.5E-4 | 443 | 516 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 1.71E-16 | 630 | 736 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-17 | 635 | 737 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.19E-13 | 636 | 734 | No hit | No description |
| PROSITE profile | PS50297 | 17.263 | 642 | 736 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 4.4E-7 | 648 | 737 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.055 | 675 | 704 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.618 | 675 | 707 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 670 | 714 | 743 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 4.2 | 848 | 870 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.718 | 849 | 878 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0056 | 850 | 869 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 2.12E-7 | 850 | 899 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.0096 | 871 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.633 | 872 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.4E-4 | 874 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1038 aa Download sequence Send to blast |
MAEVRKYGLP NQPPDISQIL LEAQNRWLRP TEICHILSNY KKFSIAPEPP NRPASGSLFL 60 FDRKILRYFR KDGHNWRKKK DGKTVKEAHE KLKVGSVDVL HCYYAHGEEN ENFQRRTYWL 120 LEEGFMNIVL VHYLEVKGGK QSFSRSKEAE ESAGLSNADS PACSNSFASQ SQVASQSMDA 180 ESPISGQISE YEDAETEKFG ATDNCRASSR YHPFVEMQQP VDGVMMNNML GVSAPRYHGE 240 MQTTTANSDN HFATHYDIAG VFNEAGAGLR GVSKTLHDSV RFAEPYPECS AEFMEPALYS 300 SNATMESNNL DDNSRLETFM SEALYTNNLT QKEADALSAA GIMSSQAENN SYTDGIRYPL 360 LKQSSLDLFK IEPDGLKKFD SFSRWMSSEL PEVADLDIKS SSDAFWSSTE TVNVADGTSI 420 PINEQLDAFA VSPSLSQDQL FSIIDVSPSY ACTGSRNKVL ITGTFLANKE HVENCKWSCM 480 FGDVEVPAEV LAHGSLRCYT PVHLSGRVPF YVTCSNRVAC SEVREFEFRD SDARQMDTSD 540 PQTTGINEMH LHIRLEKLLS LGPDDYEKYV MSDGKEKSEI INTISSLMLD DKCLNQAVPL 600 DEKEVSTARD QNIEKLVKEK LYCWLVHKVH DEDKGPNVLG KEGQGVIHLV AALGYDWAVR 660 PIITAGVKVN FRDARGWTAL HWAASCGRER TVGALIANGA ESGLLTDPTP QFPAGRTAAD 720 LASENGHKGI AGFLAESALT SHLSALTLKE SKDGNVKEIC GLGGAEDFAE SSSAQLAYRD 780 SQAESLKDSL SAVRKSTQAA ARIFQAFRVE SFHRKKVVEY GDDDCGLSDE RTLSLVSIKN 840 AKPGQNDGSH SAAVRIQNKF RGWKGRKEFM IIRQKIVKIQ AHVRGHQVRK SYRRIVWSVG 900 IVEKIILRWR RKRRGLRGFQ PVKQLEGPSP IQQLEGPSQI QPAKEEEEDE YDYLKDGRKQ 960 AEGRLQRALA RVKSMTQYPE AREQYSRIAN RVTELQEPQA MMIQDDMQSD GAIADGGDFM 1020 AELEELCGDG DAPMPTIL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
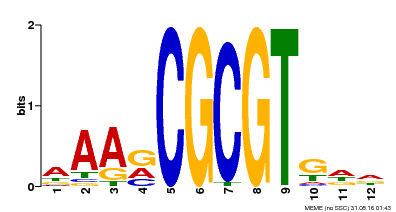
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK108461 | 0.0 | AK108461.1 Oryza sativa Japonica Group cDNA clone:002-143-D08, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015647012.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X1 | ||||
| TrEMBL | A0A0D3GTQ7 | 0.0 | A0A0D3GTQ7_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART07G22710.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.1 | 0.0 | ethylene induced calmodulin binding protein | ||||



