 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G09410.1 | ||||||||
| Common Name | CAMTA1, CMTA1, EICBP.B, SR2, T5E8.210 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 989aa MW: 111987 Da PI: 6.2294 | ||||||||
| Description | ethylene induced calmodulin binding protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 188.2 | 7.9e-59 | 22 | 139 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrc 98
l+e ++rwl+++ei++iL+n++k+++++e++trp sgsl+L++rk++ryfrkDG++w+kkkdgkt+rE+hekLKvg+++vl+cyYah+e n++fqrrc
AT5G09410.1 22 LSEaQHRWLRPTEICEILQNYHKFHIASESPTRPASGSLFLFDRKVLRYFRKDGHNWRKKKDGKTIREAHEKLKVGSIDVLHCYYAHGEANENFQRRC 119
45559********************************************************************************************* PP
CG-1 99 ywlLeeelekivlvhylevk 118
yw+Le++l +iv+vhylevk
AT5G09410.1 120 YWMLEQHLMHIVFVHYLEVK 139
*****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 84.844 | 18 | 144 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.3E-83 | 21 | 139 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.5E-51 | 24 | 138 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 1.1E-4 | 369 | 473 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.1E-4 | 394 | 469 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 7.28E-15 | 394 | 480 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF12796 | 3.3E-7 | 576 | 654 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 8.1E-18 | 577 | 689 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.624 | 577 | 688 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.51E-18 | 578 | 688 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.99E-14 | 583 | 686 | No hit | No description |
| SMART | SM00248 | 0.0029 | 627 | 656 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.22 | 627 | 659 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 8.6 | 802 | 824 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 2.02E-7 | 803 | 852 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.327 | 803 | 832 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0032 | 806 | 823 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 825 | 847 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.395 | 826 | 850 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.3E-4 | 828 | 847 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0001077 | Molecular Function | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000005 | anatomy | cultured plant cell | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009001 | anatomy | fruit | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025195 | anatomy | pollen tube cell | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001016 | developmental stage | L mature pollen stage | ||||
| PO:0001017 | developmental stage | M germinated pollen stage | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 989 aa Download sequence Send to blast |
MVDRRSFGSI TPPLQLDMEQ LLSEAQHRWL RPTEICEILQ NYHKFHIASE SPTRPASGSL 60 FLFDRKVLRY FRKDGHNWRK KKDGKTIREA HEKLKVGSID VLHCYYAHGE ANENFQRRCY 120 WMLEQHLMHI VFVHYLEVKG NRTSIGMKEN NSNSVNGTAS VNIDSTASPT STLSSLCEDA 180 DTGNRYGWTP APGMRNVSQV HGNRVRESDS QRLVDVRALD TVGNSLTRFH DQPYCNNLLT 240 QMQPSNTDSM LVEENSEKGG RLKAEHIRNP LQTQFNWQDD TDLALFEQSA QDNFETFSSL 300 LGSENLQPFG ISYQAPPSNM DSEYMPVMKI LRRSEDSLKK VDSFSKWAIK ELGEMEDLQM 360 QSSRGDIAWT TVECETAAAG ISLSPSLSED QRFTIVDFWP KSAKTDAEVE VMVIGTFLLS 420 PQEVTKYNWS CMFGEVEVPA EILVDGVLCC HAPPHTAGHV PFYVTCSNRF ACSEVREFDF 480 LSGSTQKINA TDVYGTYTNE ASLQLRFEKM LAHRDFVHEH HIFEDVGDKR RQISKIMLLK 540 EEKEYLLPGT YQRDSTKQEP KGQLFRELFE EELYIWLIHK VTEEGKGPNI LDEDGQGILH 600 FVAALGYDWA IKPVLAAGVN INFRDANGWS ALHWAAFSGR EETVAVLVSL GADAGALTDP 660 SPELPLGKTA ADLAYANGHR GISGFLAESS LTSYLEKLTV DSKENSPANS CGEKAVQTVS 720 ERTAAPMTYG DVPEKLSLKD SLTAVRNATQ AADRLHQVFR MQSFQRKQLC DIGDDEKIDI 780 SDQLAVSFAA SKTKNPGQGD VSLSCAATHI QKKYRGWKKR KEFLLIRQRI VKIQAHVRGH 840 QVRKQYRTVI WSVGLLEKII LRWRRKGNGL RGFKRNAVAK TVEPEPPVSA ICPRIPQEDE 900 YDYLKEGRKQ TEERLQKALT RVKSMVQYPE ARDQYRRLLT VVEGFRENEA SSSASINNKE 960 EEAVNCEEDD FIDIESLLND DTLMMSISP |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.26762 | 0.0 | flower| seed| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 245910_at | 0.0 | ||||
| Expression Atlas | AT5G09410 | - | ||||
| AtGenExpress | AT5G09410 | - | ||||
| ATTED-II | AT5G09410 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: During leaf developlement, expressed in young leaf primordia, hydathodes of young leaf tips, hydathodes of the lobes in maturating leaves and lamina within the minor veins in adult leaves. During flower development, expressed in developing stamens, germinating pollen grains, ovules and developing seeds. {ECO:0000269|PubMed:20383645}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, pollen and siliques. {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | calmodulin-binding protein, similar to another ethylene-upregulated calmodulin-binding protein ER1 GI:11612392 from (Nicotiana tabacum) | |||||
| UniProt | Transcription activator that binds calmodulin in a calcium-dependent manner in vitro (PubMed:11925432). Binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance (PubMed:19270186). Involved in freezing tolerance in association with CAMTA2 and CAMTA3. Contributes together with CAMTA2 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved in drought stress responses by regulating several drought-responsive genes (PubMed:23547968). Involved in auxin signaling and responses to abiotic stresses (PubMed:20383645). Activates the expression of the V-PPase proton pump AVP1 in pollen (PubMed:14581622). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:11925432, ECO:0000269|PubMed:14581622, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:23547968, ECO:0000269|PubMed:23581962, ECO:0000305|PubMed:11925432}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | 27203113 | Download |
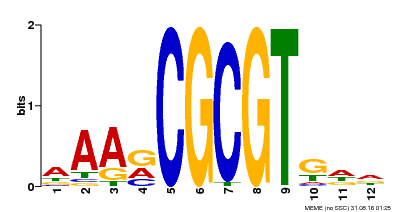
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G09410.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by UVB, wounding, ethylene and methyl jasmonate (PubMed:12218065). Induced by salt stress and heat shock (PubMed:12218065, PubMed:20383645). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G09410 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228740 | 0.0 | AK228740.1 Arabidopsis thaliana mRNA for Calmodulin-binding transcription activator 1, complete cds, clone: RAFL16-07-H14. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_196503.3 | 0.0 | ethylene induced calmodulin binding protein | ||||
| Swissprot | Q9FY74 | 0.0 | CMTA1_ARATH; Calmodulin-binding transcription activator 1 | ||||
| TrEMBL | A0A178UKF0 | 0.0 | A0A178UKF0_ARATH; EICBP.B | ||||
| STRING | AT5G09410.3 | 0.0 | (Arabidopsis thaliana) | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G09410.1 |
| Entrez Gene | 830800 |
| iHOP | AT5G09410 |
| wikigenes | AT5G09410 |



