 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016506615.1 | ||||||||
| Common Name | ABF2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 434aa MW: 46917.6 Da PI: 9.8549 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 47.6 | 3.5e-15 | 356 | 403 | 5 | 52 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleel 52
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++eN++L+k+ ee+
XP_016506615.1 356 RRQRRMIKNRESAARSRARKQAYTMELEAEVAKLKEENEELQKKQEEM 403
79***************************************9876654 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 8.8E-15 | 352 | 417 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.542 | 354 | 403 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 5.6E-15 | 356 | 405 | No hit | No description |
| Pfam | PF00170 | 3.7E-13 | 356 | 405 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 2.44E-11 | 356 | 405 | No hit | No description |
| CDD | cd14707 | 3.93E-29 | 356 | 410 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 359 | 374 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 434 aa Download sequence Send to blast |
MGSNFNFKNF GNEPPGEGGK QPGNFQLPRQ PSIYSLTFDE FLSTTGGSGK DFGSMNMDEL 60 LKNIWNAEES QTMGGSGING QEVGGVQSGH LQRQGSLTLP RTLSHKTVDE VWRDMSKEYG 120 GGKDGNSVGV PTNLPQRQQT LGEITLEEFL VRAGVVREDA QLVAKSNNVG GIFADLSYAG 180 NNTGLGFGYQ HQQTNRNTGE TGIQAANLPL NVNGVRSTQQ QLRAQQLQQQ QPQPQQQQPH 240 QQPLFPKQPA LAYAAPMAIP NSGQLGSPGM RGGMVGISDP AINSNFVQGA ALTGGGIGMV 300 GLGAGGVTVA TASPAVSSDG LGKSNGDTPS VSPLPYVFNG GIRGRKYSTV EKVVERRQRR 360 MIKNRESAAR SRARKQAYTM ELEAEVAKLK EENEELQKKQ EEMLEIQKNQ VMEMMNLQKG 420 AKRRCLRRTQ TGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in ABA and stress responses and acts as a positive component of glucose signal transduction. Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
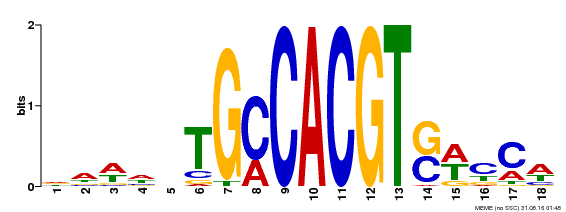
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA), cold and glucose. {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF736850 | 0.0 | KF736850.1 Nicotiana tabacum ABRE binding factor (ABF2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001313058.1 | 0.0 | bZIP transcription factor TRAB1-like | ||||
| Refseq | XP_016506615.1 | 0.0 | PREDICTED: bZIP transcription factor TRAB1-like | ||||
| Swissprot | Q9M7Q4 | 1e-129 | AI5L5_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| TrEMBL | V9XND3 | 0.0 | V9XND3_TOBAC; ABRE binding factor | ||||
| STRING | XP_009592272.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA776 | 24 | 97 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G45249.1 | 2e-98 | abscisic acid responsive elements-binding factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 107824387 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



