 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016451280.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 302aa MW: 33894 Da PI: 9.3571 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.5 | 5.5e-32 | 119 | 172 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt++LH++Fv+av+ LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
XP_016451280.1 119 PRMRWTSTLHAHFVHAVQLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 172
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.68E-15 | 116 | 173 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.7E-28 | 117 | 173 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.7E-24 | 119 | 172 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.8E-7 | 120 | 171 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 302 aa Download sequence Send to blast |
MRDTISSSMP DLSLQISLPS NILKCDNHSF STESGSSGGS DLSHENNGIF RPQKAFNNLP 60 LLTSFEPTLS LGFGQLPNRN LQRYHQHQHH QQYQPQIYGR DFKRSSRMIS GVKRSVRAPR 120 MRWTSTLHAH FVHAVQLLGG HERATPKSVL ELMNVKDLTL AHVKSHLQMY RTVKSTDKTT 180 AKGMTEIFVN PNRGNNISEY EKSCEEIDAM TDPSVSLSNT IPQNAHIRIP SSFMTETNAW 240 SHSNRERAFP YPHIYSNMKV IEPADNSTSS DTLVNLEFTL GRPSLNMGHA GSSSNDSTLL 300 KC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-17 | 120 | 174 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
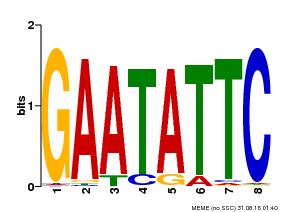
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KP271989 | 3e-74 | KP271989.1 UNVERIFIED: Populus trichocarpa isolate Potri.014G037200_BESC-125 mRNA sequence. | |||
| GenBank | KP271990 | 3e-74 | KP271990.1 UNVERIFIED: Populus trichocarpa isolate Potri.014G037200_BESC-293 mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009804554.1 | 0.0 | PREDICTED: probable transcription factor KAN4 isoform X2 | ||||
| Refseq | XP_016451280.1 | 0.0 | PREDICTED: probable transcription factor KAN4 | ||||
| TrEMBL | A0A1S3YGM7 | 0.0 | A0A1S3YGM7_TOBAC; probable transcription factor KAN4 | ||||
| TrEMBL | A0A1U7YZK0 | 0.0 | A0A1U7YZK0_NICSY; probable transcription factor KAN4 isoform X2 | ||||
| STRING | XP_009804553.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12262 | 18 | 22 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.1 | 8e-63 | G2-like family protein | ||||



