 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | NNU_009511-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; stem eudicotyledons; Proteales; Nelumbonaceae; Nelumbo
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 265aa MW: 30219.9 Da PI: 6.6232 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47 | 5.8e-15 | 10 | 64 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHT.......TTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmg.......kgRtlkqcksrwqkyl 48
+g+WT++Ed++lv+ v ++G ++W+ Ia+ g + Rt+k+c++rw +yl
NNU_009511-RA 10 KGPWTEQEDLQLVCFVGLFGERRWDFIAKVSGskvagdtTARTGKSCRLRWVNYL 64
79***************************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 56.4 | 7e-18 | 70 | 115 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgr+T++E++l ++++ ++G++ W++Iar+++ gRt++++k++w++++
NNU_009511-RA 70 RGRMTPQEEQLVLELHSRWGNR-WSRIARKLP-GRTDNEIKNYWRTHM 115
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.3 | 5 | 64 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.91E-27 | 8 | 111 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.7E-13 | 9 | 66 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.4E-14 | 10 | 64 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-18 | 11 | 71 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.65E-10 | 12 | 64 | No hit | No description |
| PROSITE profile | PS51294 | 26.583 | 65 | 119 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.3E-16 | 69 | 117 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.0E-16 | 70 | 114 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.38E-11 | 72 | 115 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 5.8E-24 | 72 | 118 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 265 aa Download sequence Send to blast |
MKMVQEEIRK GPWTEQEDLQ LVCFVGLFGE RRWDFIAKVS GSKVAGDTTA RTGKSCRLRW 60 VNYLHPGLKR GRMTPQEEQL VLELHSRWGN RWSRIARKLP GRTDNEIKNY WRTHMRKKAQ 120 ERKRSISPSS SSSQCSSPST NNPPAASSIS LPETHNGEGI SSLHNCGSST DLSAVRGEVS 180 EEADAIWYPM DDQIWKEFPS VEEAISGPVY EGRKDEGCSS YPPMASPLWD YCCSDTLWRM 240 DEEEIKMILP MADHFVSTYG HDTRE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 1e-27 | 7 | 119 | 1 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| 1h88_C | 2e-27 | 2 | 119 | 50 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 2e-27 | 2 | 119 | 50 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00600 | PBM | Transfer from AT5G59780 | Download |
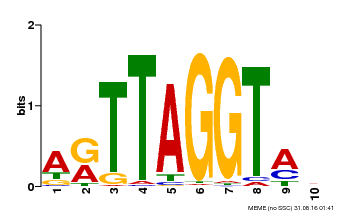
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Isoform MYB59-1 is induced by jasmonate (JA), salicylic acid (SA), gibberellic acid (GA), and ethylene. Also induced by cadmium (Cd). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:16531467}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010260059.1 | 0.0 | PREDICTED: transcription factor MYB59-like isoform X1 | ||||
| Swissprot | Q4JL84 | 3e-80 | MYB59_ARATH; Transcription factor MYB59 | ||||
| TrEMBL | A0A1U7ZZN5 | 0.0 | A0A1U7ZZN5_NELNU; transcription factor MYB59-like isoform X1 | ||||
| STRING | XP_010260059.1 | 0.0 | (Nelumbo nucifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G59780.3 | 3e-80 | myb domain protein 59 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



