 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Niben101Scf22883g00002.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 342aa MW: 38909.3 Da PI: 4.3386 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.3 | 6.3e-19 | 53 | 106 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
Niben101Scf22883g00002.1 53 KKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 106
56689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.2 | 8.5e-42 | 52 | 144 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeve 85
ekkrrls eqvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy +Lk+++dalk++ e+L++++e
Niben101Scf22883g00002.1 52 EKKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGVLKANFDALKHNYESLKHDNE 136
69*********************************************************************************** PP
HD-ZIP_I/II 86 eLreelke 93
+L +e++e
Niben101Scf22883g00002.1 137 ALLKEIRE 144
**999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.88E-19 | 43 | 110 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.491 | 48 | 108 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.1E-18 | 51 | 112 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.03E-16 | 53 | 109 | No hit | No description |
| Pfam | PF00046 | 4.1E-16 | 53 | 106 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 4.6E-20 | 55 | 115 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.0E-5 | 79 | 88 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 83 | 106 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.0E-5 | 88 | 104 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 3.0E-18 | 108 | 148 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 342 aa Download sequence Send to blast |
MKRPCSSDSL GALMSMCPNT TDEQNHVYST RDFQSMLLDE EGCIEESGHI SEKKRRLSVE 60 QVKALEKNFE VENKLEPERK VKLAQELGLQ PRQVAVWFQN RRARWKTKQL ERDYGVLKAN 120 FDALKHNYES LKHDNEALLK EIRELKSKVY TGNGESKGVA VKEEAMESES DDNKLIEQSK 180 PNDNDNNNNN NFLQNFEEED DEEEEINFEN FNVAAASTNI FGDNFKDGSS DSDSSAILNE 240 DNSPNAAAIS SSGAFLISTN NGNGKGSSTS LNFCFQFTES SSKSNLGDGQ KGDNYYYQPQ 300 QYVKMEEHNF FNGEESCSTL FTDEQAPTLQ WYCPEDWNWK ES |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 100 | 108 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
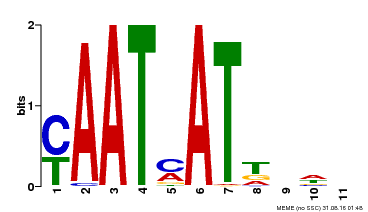
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009777009.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-6-like | ||||
| Refseq | XP_016502437.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-6-like | ||||
| TrEMBL | A0A1S4CMF7 | 0.0 | A0A1S4CMF7_TOBAC; homeobox-leucine zipper protein ATHB-6-like | ||||
| TrEMBL | A0A1U7WQP6 | 0.0 | A0A1U7WQP6_NICSY; homeobox-leucine zipper protein ATHB-6-like | ||||
| STRING | XP_009777009.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2718 | 24 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 1e-65 | homeobox protein 6 | ||||



