 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Niben101Scf15224g00002.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 643aa MW: 70077.8 Da PI: 5.2572 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 35.2 | 2.2e-11 | 471 | 516 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ +I++L
Niben101Scf15224g00002.1 471 NHVEAERQRREKLNQRFYALRAVVPNV-----SKMDKASLLGDAIAFINEL 516
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.2E-53 | 67 | 244 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.539 | 467 | 516 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 5.34E-14 | 470 | 521 | No hit | No description |
| SuperFamily | SSF47459 | 1.7E-17 | 470 | 540 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 6.3E-9 | 471 | 516 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 5.7E-18 | 471 | 537 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 2.0E-15 | 473 | 522 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009269 | Biological Process | response to desiccation | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009867 | Biological Process | jasmonic acid mediated signaling pathway | ||||
| GO:0009963 | Biological Process | positive regulation of flavonoid biosynthetic process | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0043619 | Biological Process | regulation of transcription from RNA polymerase II promoter in response to oxidative stress | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051090 | Biological Process | regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:2000068 | Biological Process | regulation of defense response to insect | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 643 aa Download sequence Send to blast |
MTNIWSNTTS DDNMMEAFLS SDPSSFWPGT TTTTPTPRSS VSPAPAPVTG IAVDPLTSMP 60 YFNQESLQQR LQTLIDGARE AWTYAIFWQS PSVLGWGDGY YKGEEDKNKR KTASLSPDFI 120 TEQAHRKKVL RELNSLISGT QTGGENDAVD EEVTDTEWFF LISMTQSFVN GSGLPGLAMY 180 SSSPIWVTGA ERLAASHCER ARQAQGFGLQ TIVCIPSANG VVELGSTELI FQTADLMNKV 240 KVLFNFNIDM GATTGSGSGS CAIQAEPDPS ALWLTDPASS VAEVKDSANT IPSSNSSKQL 300 VFGNENSENG HQNSQQTQGF FTRELNFSEY GFDGSNTRNG NANSSRSCKP ESGEILNFGD 360 STKRSVSSAN GSLFSGQSQF GAGPAEENKN KNKKRSSASR GSNDEGMLSF VSGVILPSSN 420 TGKSSGGGDS DQSDLEASVV KEADSSRVVD PEKKPRKRGR KPANGREEPL NHVEAERQRR 480 EKLNQRFYAL RAVVPNVSKM DKASLLGDAI AFINELKSKV QNSDSDKEEL RNQIESLRKE 540 LANKGSNYTG PPPSNQELKI VDMDIDVKVI GWDAMIRIQS NKKNHPAARL MAALMELDLD 600 VHHASVSVVN ELMIQQATVK MESRLYTQEQ LRISLTSRIA ESR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 6e-65 | 60 | 247 | 2 | 195 | Transcription factor MYC3 |
| 4rqw_B | 6e-65 | 60 | 247 | 2 | 195 | Transcription factor MYC3 |
| 4rs9_A | 6e-65 | 60 | 247 | 2 | 195 | Transcription factor MYC3 |
| 4yz6_A | 6e-65 | 60 | 247 | 2 | 195 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 452 | 460 | KKPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds to the G-box motif (5'-AACGTG-3') found in a number of promoters of jasmonate-induced genes (PubMed:15231736). Transcription activator involved in the transcriptional regulation of terpene biosynthesis in glandular trichomes (PubMed:24884371, PubMed:30518626). Binds to the promoter of the linalool synthase TPS5 and promotes TPS5 gene transactivation (PubMed:24884371). Acts synergistically with EOT1 in the transactivation of TPS5 (PubMed:24884371). Involved in type VI glandular trichome development (PubMed:30518626). Involved in the activation of terpene synthases required for volatile mono- and sesquiterpenes synthesis by the glandular cells of type VI trichomes (PubMed:30518626). {ECO:0000269|PubMed:15231736, ECO:0000269|PubMed:24884371, ECO:0000269|PubMed:30518626}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00084 | PBM | Transfer from AT1G32640 | Download |
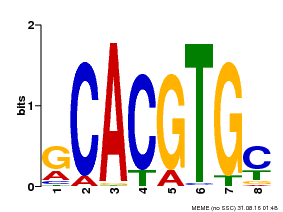
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by methyl jasmonate and wounding. {ECO:0000269|PubMed:15231736}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009800420.1 | 0.0 | PREDICTED: transcription factor MYC2-like | ||||
| Swissprot | A0A060KY90 | 0.0 | MYC1_SOLLC; Transcription factor MYC1 | ||||
| TrEMBL | D7P229 | 0.0 | D7P229_NICBE; BHLH2 transcription factor | ||||
| STRING | XP_009800420.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3300 | 24 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G32640.1 | 0.0 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



