 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Niben101Scf02853g02002.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1241aa MW: 138845 Da PI: 9.1608 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.2 | 0.00055 | 1208 | 1234 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
Niben101Scf02853g02002.1 1208 YVCTetGCGQTFRFVSDFSRHKRKtgH 1234
99********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 3.6E-16 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.991 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.1E-15 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 1.9E-49 | 180 | 349 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 31.421 | 180 | 349 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 4.67E-24 | 194 | 338 | No hit | No description |
| Pfam | PF02373 | 2.4E-37 | 213 | 332 | IPR003347 | JmjC domain |
| SMART | SM00355 | 24 | 1125 | 1147 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.468 | 1148 | 1177 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.015 | 1148 | 1172 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.3E-5 | 1149 | 1176 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1150 | 1172 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.89E-9 | 1164 | 1204 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.8E-8 | 1177 | 1202 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1178 | 1207 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1178 | 1202 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1180 | 1202 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.86E-8 | 1196 | 1230 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-9 | 1203 | 1231 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.19 | 1208 | 1234 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.053 | 1208 | 1239 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1210 | 1234 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1241 aa Download sequence Send to blast |
MAEASGNIEV FPWLKTLPVA PEYHPTLEEF QDPIAYIFKI EKEASKYGIC KIVPPVPAPP 60 KKTALSYLNR SLSARSGSNG GPTFTTRQQQ IGFCPRKHRP VKKPVWQSGE NYTVQEFQAK 120 AKAFEKNFLK KNLKKALTSP LEIETLYWKA TVDKPFCVEY ANDMPGSAFA PKKVGNAVTT 180 LADTEWNMRG VSRAKGSLLK FMKEEIPGVT SPMVYLAMMF SWFAWHVEDH DLHSLNYLHM 240 GAGKTWYGVP RDAAVAFEEV IRVHGYAGET NPLVTFATLG EKTTVMSPEV LLSAGIPCCR 300 LVQNAGEFVI TFPRAYHSGF SHGYNCGEAS NIATPGWLIV AKDAAIRRAS INCPPMVSHF 360 QLLYDLALSL CSRVPKNIRI EPRSSRLKDK KKSEGDMLVK ELFVEDLNCN NYLLHILGEG 420 SPVVLLPRHY SGISIGSNSV AGSQLKVNSR FPSLSSPDHE VKSKTDSASD ALMLGRKLRT 480 KQLASVSLEK GKHSSWHAGN RLPESGRDEA ESSPETEGGN LDSARGMTYR CDTLSEHGLF 540 SCVTCGILCY TCVAIIQPTE TAAHHLMSSD YRNFNDWTGN VGGITATSRD ANAAESDSSS 600 GMLVKRAPGG LIDVPIESSD RIRKLNNERV GVLSSTKARK ETSSLGLLAL NYANSSDSDE 660 DEVEANIPVE TCESRHMDFE DEVSLRVIDP YANHRQRRAV SDGRNCQSRD NSIHLESENP 720 PSGESNTLPD RARHQPRSHQ VGANCIPFSH RGEIANSDVV VPFDNGPMQF TSTSDEDSFR 780 IHVFCLQHAV QVEEQLRQVG GVHISLLCHP DYPKLEAQAK KVVEELGRDH FWREISFREA 840 TKEDEEMIQS ALEVEDAIHG NGDWTVKLDI NLFYSSNLSR SPLYSKQMPY NFIIYNAFGR 900 SSPDEKSEYT GTGRGSGKQK RAVVAGKWCG KIWMSSQVHP LLAERRDTDE EQQLNNNSIS 960 ARVKPEVKSE RPCEMTQTIK TVARTGKKRE SRVENKRKSK LLVADDSSVS NVPQQQRKTN 1020 LRSKRIKYET PEPKDEDVDK KKRFDSPIND DPVGGPSTRL RKRMPKPSSE SLVKPRPTPI 1080 KQQNESKKAE KGSKVKIPSA NSNSKKDPVM KANTSKKMKD KEGEYHCDLE GCSMSFNAKQ 1140 ELSLHKKNIC PVEGCKKKFF SHKYLVQHRR VHMDDRPLKC PWKGCKMTFK WAWARTEHIR 1200 VHTGARPYVC TETGCGQTFR FVSDFSRHKR KTGHVSKKGR G |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 5e-71 | 13 | 382 | 8 | 366 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 5e-71 | 13 | 382 | 8 | 366 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
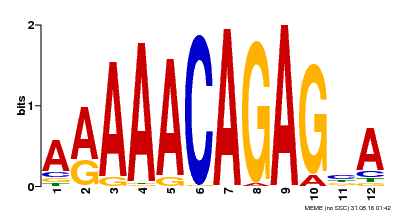
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019243504.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A1J6IKD2 | 0.0 | A0A1J6IKD2_NICAT; Lysine-specific demethylase ref6 | ||||
| STRING | XP_009613168.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4988 | 23 | 31 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



