 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr8g090205.7 | ||||||||
| Common Name | MTR_8g090205 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 950aa MW: 107664 Da PI: 5.788 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 175.7 | 6.1e-55 | 18 | 134 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
l+ + rwl++ ei++iL+n++++++++++ p+sgsl+L++rk +ryfrkDG++w+kkkdgktvrE+he+LK g+vevl+cyYah+e+n +fq
Medtr8g090205.7 18 LQAQYRWLRPAEICQILTNYNSFQISSQPSYMPPSGSLFLFDRKATRYFRKDGHNWRKKKDGKTVREAHERLKAGSVEVLHCYYAHGEQNDNFQ 111
67799***************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Leeel++ivlvhy+evk
Medtr8g090205.7 112 RRTYWMLEEELSHIVLVHYREVK 134
********************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 76.211 | 13 | 139 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.6E-76 | 16 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.4E-49 | 20 | 132 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.27E-15 | 507 | 592 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 4.5E-4 | 507 | 583 | IPR002909 | IPT domain |
| Gene3D | G3DSA:1.25.40.20 | 3.7E-17 | 690 | 804 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.6 | 693 | 804 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.87E-17 | 699 | 804 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.54E-13 | 699 | 802 | No hit | No description |
| Pfam | PF12796 | 1.1E-7 | 715 | 805 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.11 | 743 | 772 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 500 | 782 | 811 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50096 | 6.852 | 911 | 939 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 950 aa Download sequence Send to blast |
MSNSYTMMAT TDIEEILLQA QYRWLRPAEI CQILTNYNSF QISSQPSYMP PSGSLFLFDR 60 KATRYFRKDG HNWRKKKDGK TVREAHERLK AGSVEVLHCY YAHGEQNDNF QRRTYWMLEE 120 ELSHIVLVHY REVKRTKATL IHAKENEESN PCDQQSYKVM PNTEAETSLP SSMNSGQASE 180 YEEAESAFNS HANSDFYSFL ELQQPAVQKI KAQLPYSNCP LPLKDDQERL PVIPQVDDIS 240 LSQTNETKYI NNVGLTCELS KVLGFSSWQD ILENKAGSHN VPFQSFPEKE PNNMEINSTS 300 QGYETMGQHL TISITKQHEN RSFIQAEAEG NWQASGFNSL SASTCPKDSA YSGSSCEVTY 360 SDNEQEVNEV DFQQSLEQFL LHAHQQHKVC MRNSSHEIPL KAEDRLKSDL GVDKSPDGIE 420 DTQFTSKKTI LSVSVAEDGL KKLDSFNQWM SKELCDVEES SKHSPSGAYW DTVESENGVD 480 STTIPSQVHL ENYVLDPSIC YDQLFSIIDY SPSWTFEDSE IEVLISGRFL KSQHEAEDCK 540 WSCMFGEIEV PAEITRNGVL CCHTPQHKAG RVPFYVTCSN RLACSEVREF DFRVNYTQED 600 NTAGETRSRN TYDTFNKRFG EFLSQEHDFP RVLDSISVNE KSQLRSKIGS LLGRKDDEWD 660 ELLKFTLDKD FSPELVHDQL LQNLLKDKLH AWLLQKTTED GKGPNVLDEG GQGVLHFAAF 720 FGYDWAFEPT IVAGVNVNFR DVNGWTALHW AAFCGRERTV ASLISLGGAP GALTDPCPQH 780 PSGRTPADLA SANGHKGIAG YLAESFLSSQ LKSLDLKRNM RETVGTKQRV QEQNNECFSH 840 EPSMKDSLAA VCNATQAAAR IHQVFRVQSF QRKQQKEYDG DKFGISDERA LSLITVNAKS 900 HKSGLRIEPV HVAATRIQNK FRSWKGRKDF LIIRQRIVKI QVHIVPTFA* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.20634 | 2e-44 | root | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
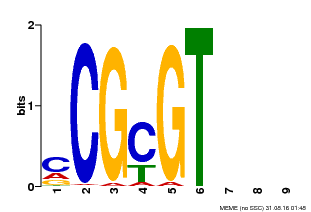
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr8g090205.7 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013446800.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X1 | ||||
| TrEMBL | A0A072TVD4 | 0.0 | A0A072TVD4_MEDTR; Calmodulin-binding transcription activator | ||||
| STRING | XP_004504077.1 | 0.0 | (Cicer arietinum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr8g090205.7 |
| Entrez Gene | 25501911 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



