 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr8g090205.2 | ||||||||
| Common Name | MTR_8g090205 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1085aa MW: 123480 Da PI: 6.1469 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 175.4 | 7.3e-55 | 16 | 132 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
l+ + rwl++ ei++iL+n++++++++++ p+sgsl+L++rk +ryfrkDG++w+kkkdgktvrE+he+LK g+vevl+cyYah+e+n +fq
Medtr8g090205.2 16 LQAQYRWLRPAEICQILTNYNSFQISSQPSYMPPSGSLFLFDRKATRYFRKDGHNWRKKKDGKTVREAHERLKAGSVEVLHCYYAHGEQNDNFQ 109
67799***************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Leeel++ivlvhy+evk
Medtr8g090205.2 110 RRTYWMLEEELSHIVLVHYREVK 132
********************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 76.211 | 11 | 137 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.6E-76 | 14 | 132 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.8E-49 | 18 | 130 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.73E-15 | 505 | 590 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.3E-4 | 505 | 581 | IPR002909 | IPT domain |
| Gene3D | G3DSA:1.25.40.20 | 4.7E-17 | 688 | 802 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.6 | 691 | 802 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.18E-17 | 697 | 802 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.06E-13 | 697 | 800 | No hit | No description |
| Pfam | PF12796 | 1.3E-7 | 713 | 803 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.11 | 741 | 770 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 500 | 780 | 809 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.65E-7 | 902 | 958 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 22 | 907 | 929 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 6.852 | 909 | 937 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.14 | 910 | 928 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0039 | 930 | 952 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.597 | 931 | 955 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 933 | 952 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1085 aa Download sequence Send to blast |
MAHLHPNQFD IEEILLQAQY RWLRPAEICQ ILTNYNSFQI SSQPSYMPPS GSLFLFDRKA 60 TRYFRKDGHN WRKKKDGKTV REAHERLKAG SVEVLHCYYA HGEQNDNFQR RTYWMLEEEL 120 SHIVLVHYRE VKRTKATLIH AKENEESNPC DQQSYKVMPN TEAETSLPSS MNSGQASEYE 180 EAESAFNSHA NSDFYSFLEL QQPAVQKIKA QLPYSNCPLP LKDDQERLPV IPQVDDISLS 240 QTNETKYINN VGLTCELSKV LGFSSWQDIL ENKAGSHNVP FQSFPEKEPN NMEINSTSQG 300 YETMGQHLTI SITKQHENRS FIQAEAEGNW QASGFNSLSA STCPKDSAYS GSSCEVTYSD 360 NEQEVNEVDF QQSLEQFLLH AHQQHKVCMR NSSHEIPLKA EDRLKSDLGV DKSPDGIEDT 420 QFTSKKTILS VSVAEDGLKK LDSFNQWMSK ELCDVEESSK HSPSGAYWDT VESENGVDST 480 TIPSQVHLEN YVLDPSICYD QLFSIIDYSP SWTFEDSEIE VLISGRFLKS QHEAEDCKWS 540 CMFGEIEVPA EITRNGVLCC HTPQHKAGRV PFYVTCSNRL ACSEVREFDF RVNYTQEDNT 600 AGETRSRNTY DTFNKRFGEF LSQEHDFPRV LDSISVNEKS QLRSKIGSLL GRKDDEWDEL 660 LKFTLDKDFS PELVHDQLLQ NLLKDKLHAW LLQKTTEDGK GPNVLDEGGQ GVLHFAAFFG 720 YDWAFEPTIV AGVNVNFRDV NGWTALHWAA FCGRERTVAS LISLGGAPGA LTDPCPQHPS 780 GRTPADLASA NGHKGIAGYL AESFLSSQLK SLDLKRNMRE TVGTKQRVQE QNNECFSHEP 840 SMKDSLAAVC NATQAAARIH QVFRVQSFQR KQQKEYDGDK FGISDERALS LITVNAKSHK 900 SGLRIEPVHV AATRIQNKFR SWKGRKDFLI IRQRIVKIQA HVRGHQVRKN YRKIIWSVGI 960 VEKIILRWRR KGSGLRGFKS EAISEGTMVQ GVSSATEDDY DFLKEGRKQT EKRLEKALAR 1020 VKSMAQYPDA RDQYHRLLNV VTEIQENQVK QDRNFINSEE SRQFNSLVDL EALLDEDTFM 1080 PTAT* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.20634 | 0.0 | root | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
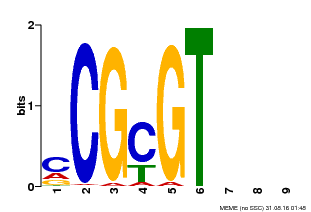
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr8g090205.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013446801.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X2 | ||||
| TrEMBL | A0A072TVC9 | 0.0 | A0A072TVC9_MEDTR; Calmodulin-binding transcription activator | ||||
| STRING | XP_004504077.1 | 0.0 | (Cicer arietinum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr8g090205.2 |
| Entrez Gene | 25501911 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



